Tutorial guidelines¶
Steps in GUI tutorials¶
For steps in tutorials use the rst-class:: steps.
Example:
.. rst-class:: steps
| Optionally some context here
| **1.** The first step, e.g. click on **Edit → Builder**
| **2.** The second step.
Result:
GUI icons¶
In the Tutorials, you can refer to some GUI buttons/icons by using the following substitutions (these substitutions are defined in global_conf.py, but that you have to initialize it in the conf.py of your specific project. See the Tutorials/conf.py).
Icon |
Substitution |
Name |
|---|---|---|
|
|InfoBtn| |
More-info button |
|
|MoreBtn| |
Details |
|
|SCMMenu| |
SCM menu |
|
|Search| |
Search box |
|
|PointerTool| |
Pointer tool |
|
|CTool| |
Carbon tool |
|
|OTool| |
Oxygen tool |
|
|HTool| |
Hydrogen tool |
|
|XTool| |
Element tool |
|
|StructTool| |
Structure tool |
|
|CrystalTool| |
Periodic structure tool |
|
|SymmTool| |
Symmetrize tool |
|
|BondTool| |
Bond tool |
|
|PeriodicViewTool| |
Periodic display |
|
|SliceTool| |
Slice tool |
|
|PreOptimTool| |
Pre-optimizer button |
|
|ADFPanel| |
ADF panel |
|
|ADFviaAMSPanel| |
ADF via AMS panel |
|
|BANDPanel| |
BAND panel |
|
|ConformersPanel| |
Conformers panel |
|
|DFTBPanel| |
DFTB panel |
|
|MMPanel| |
MM panel |
|
|MopacPanel| |
Mopac panel |
|
|QMMMPanel| |
QMMM panel |
|
|QuantumESPRESSOPanel| |
Quantum ESPRESSO panel |
|
|QuildPanel| |
Quild panel |
|
|ReaxFFPanel| |
ReaxFF panel |
|
|ForceFieldPanel| |
ForceField panel |
|
|VASPPanel| |
VASP panel |
|
|AddButton| |
Add button |
|
|DeleteButton| |
Delete button |
Example:
.. rst-class:: steps
| In **ADFInput**
| **1.** Switch to the **BAND panel**: |ADFPanel| **→** |BANDPanel|
| **2.** In the **search box** |Search| search for ``Diamond`` and select **Crystal → C**
Result:
 →
→ 
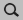 search for
search for Diamond and select Crystal → CConventions for step descriptions¶
How to phrase common tasks:
 →
→ 
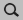 search for
search for Diamond and select Crystal → C-1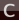 and click in the molecule drawing area
and click in the molecule drawing areaScripting Examples and screx¶
The preferred way to create new tutorials for PLAMS/Python is through Jupyter notebooks.
These are located in the $AMSHOME/userdoc/PythonExamples directory.
Important
NEVER MODIFY THE EXAMPLE index.rst FILES MANUALLY! They are generated by screx either manually or at build time.
The scripting examples use
screx.tagsto filter on tags, andscrex.notebooksto combine jupyter notebooks (.ipynb) with some metadata (conf.toml) into rst files read by sphinx. Note that the tags are defined in conf.toml!
screx is both a sphinx plugin and a command-line tool.
Recommendation: Use screx as a command-line tool to manually convert the Jupyter notebooks into .rst. The screx command-line tool has lots of functionality.
git clone [email protected]:hellstrom/screx.git
amspython -m pip install -e screx
Then you can do e.g. amspython -m screx convert my-example. See the screx docs for more detail.
The
screx.tomldefines the tag categoriesThe (jinja2) templates are in the
screx_templates/directory. These define how to render the data from conf.toml into index.rst. Screx inserts everything from the[page]into the template context, and additionally some extra info (like resolved paths for toc figures). Be careful when editing the preamble.j2 and postamble.j2 templates, since the .rst format is sensitive to newlines and indentation.screx_templates/also has thetemplate_conf.tomlfile that is the defaultconf.tomlfile when runningscrex new some-new-example.
Older way for legacy PLAMS examples¶
From these notebooks, documentation pages can be generated by running a script generate_example.sh, located in the examples directory.
This will create .rst files which can be included in the PLAMS Examples.
For full instructions on how to do this, see $AMSHOME/scripting/scm/plams/examples/README.md.