The Ziff-Gulari-Barshad (ZGB) Model¶
In the previous tutorial, we have shown how the Zacros input files can be translated to pyZacros scripts. Here, we will show some additional examples for post-processing of the simulation output.
The example script can be downloaded through this link: ZiffGulariBarshad.py.
pyZacros Setup¶
pyZacros scripting can be used to immediately visualize the output of your simulation, enabling automated report generation. Various built-in functions are available to access commonly-used reports.
The basic structure of the script is much the same as in the preceding tutorial. We start by defining the species, lattice, reactions and simulation settings. The parameters used for the Ziff-Gulari-Barshad model can be found in the models overview.
We use the run() call to start the Zacros simulation. A results object is generated once the simulation completes. This results object gives us access to the simulation output within Python. For this example, we add a check to make sure that the job has finished without errors (job.ok) before proceeding with the analysis.
scm.pyzacros.init()
results = job.run()
if job.ok():
# Post-processing & visualization
results.plot_lattice_states(results.lattice_states())
scm.pyzacros.finish()
Using the plot_lattice_states function allows us to generate snapshots showing the transient evolution of the lattice during the kMC simulation.
Running the Simulation¶
We will now run the example script using Python. For default AMS installations:
$AMSBIN/amspython WaterGasShiftOnPt111.py
An overview of the simulation settings will be shown and the jobs starts running. Once the simulation has completed, a movie will play showing the transient evolution of the lattice.
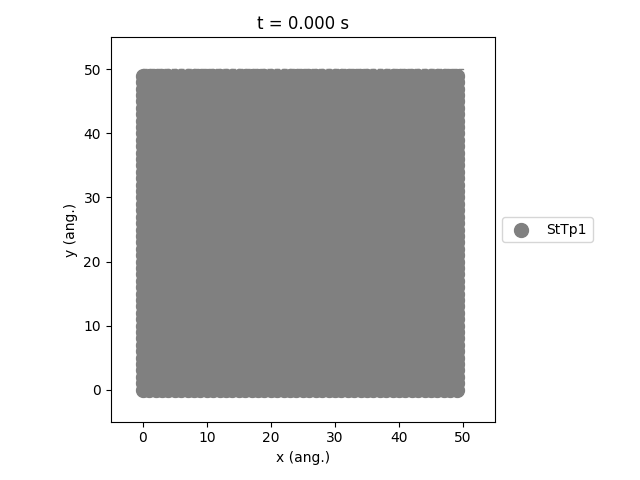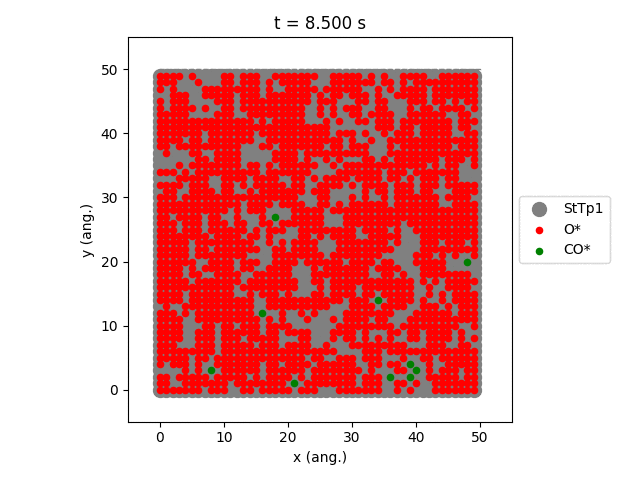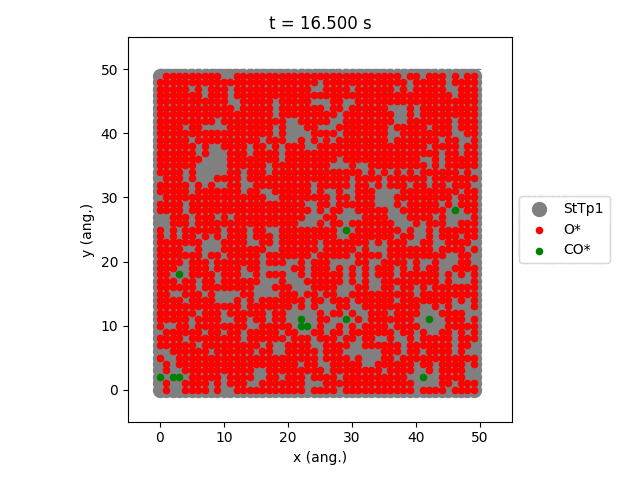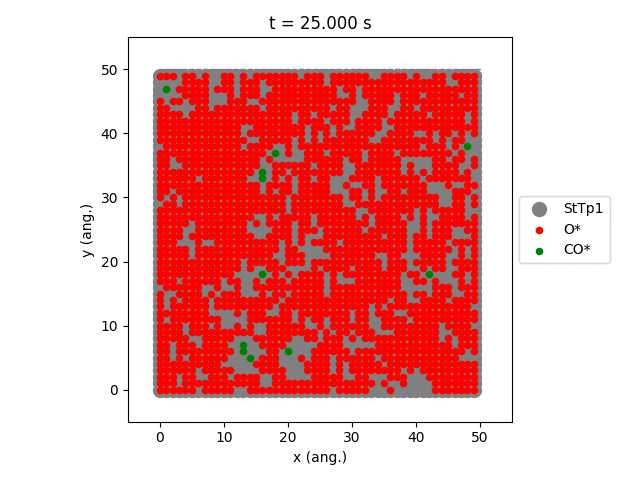
The StTp is the default name for a SiteType in Zacros. When constructing your own lattices, you can provide custom labels for the different sites, which will then also update the legend shown in the graph.
1$ amspython ZiffGulariBarshad.py
2[14.02|17:20:01] PLAMS working folder: /home/user/pyzacros/examples/ZiffGulariBarshad/plams_workdir
3---------------------------------------------------------------------
4simulation_input.dat
5---------------------------------------------------------------------
6random_seed 953129
7temperature 500.0
8pressure 1.0
9
10snapshots on time 0.5
11process_statistics on time 0.01
12species_numbers on time 0.01
13max_time 25.0
14
15n_gas_species 3
16gas_specs_names CO O2 CO2
17gas_energies 0.00000e+00 0.00000e+00 -2.33700e+00
18gas_molec_weights 2.79949e+01 3.19898e+01 4.39898e+01
19gas_molar_fracs 4.50000e-01 5.50000e-01 0.00000e+00
20
21n_surf_species 2
22surf_specs_names CO* O*
23surf_specs_dent 1 1
24
25finish
26---------------------------------------------------------------------
27lattice_input.dat
28---------------------------------------------------------------------
29lattice default_choice
30 rectangular_periodic 1.0 50 50
31end_lattice
32---------------------------------------------------------------------
33energetics_input.dat
34---------------------------------------------------------------------
35energetics
36
37cluster CO*-0
38 sites 1
39 lattice_state
40 1 CO* 1
41 site_types 1
42 graph_multiplicity 1
43 cluster_eng -1.30000e+00
44end_cluster
45
46cluster O*-0
47 sites 1
48 lattice_state
49 1 O* 1
50 site_types 1
51 graph_multiplicity 1
52 cluster_eng -2.30000e+00
53end_cluster
54
55end_energetics
56---------------------------------------------------------------------
57mechanism_input.dat
58---------------------------------------------------------------------
59mechanism
60
61step *-0:CO-->CO*-0
62 gas_reacs_prods CO -1
63 sites 1
64 initial
65 1 * 1
66 final
67 1 CO* 1
68 site_types 1
69 pre_expon 1.00000e+01
70 activ_eng 0.00000e+00
71end_step
72
73step *_0-0,*_1-0:O2-->O*_0-0,O*_1-0;(0,1)
74 gas_reacs_prods O2 -1
75 sites 2
76 neighboring 1-2
77 initial
78 1 * 1
79 2 * 1
80 final
81 1 O* 1
82 2 O* 1
83 site_types 1 1
84 pre_expon 2.50000e+00
85 activ_eng 0.00000e+00
86end_step
87
88step CO*_0-0,O*_1-0-->*_0-0,*_1-0:CO2;(0,1)
89 gas_reacs_prods CO2 1
90 sites 2
91 neighboring 1-2
92 initial
93 1 CO* 1
94 2 O* 1
95 final
96 1 * 1
97 2 * 1
98 site_types 1 1
99 pre_expon 1.00000e+20
100 activ_eng 0.00000e+00
101end_step
102
103end_mechanism
104[14.02|17:29:40] JOB plamsjob STARTED
105[14.02|17:29:40] JOB plamsjob RUNNING
106[14.02|17:29:41] JOB plamsjob FINISHED
107[14.02|17:29:41] JOB plamsjob SUCCESSFUL
108[14.02|17:32:01] PLAMS run finished. Goodbye
Post-processing¶
In order to customize your own reports, you can simply add additional function calls to the post-processing block.
if job.ok():
# Post-processing & visualization
results.plot_molecule_numbers(["CO*", "O*"])
results.plot_lattice_states(results.lattice_states())
Here, you can use any of the built-in visualization tools provided by pyZacros, or you can make your own analysis scripts by accessing the results data. (Examples for this are shown in the intermediate tutorials.)
pyZacros can also be used to load results from past jobs. This allows you to modify the visualization script without having to re-run the simulation.
job = pz.ZacrosJob.load_external( path="plams_workdir/plamsjob" )
job.results.plot_lattice_states(job.results.lattice_states())