Atomic Layer Etching of SiO2¶
This tutorial will teach you how to:
relax a silica slab with ReaxFF,
define and configure an HF projectile, and
run and inspect a molecule-gun molecular dynamics.
This tutorial is the GUI companion to AtomicLayerEtching.ipynb. The notebook builds the slab programmatically. In the workflow below, we use the prepared structure files so that the tutorial remains reproducible.
Introduction¶
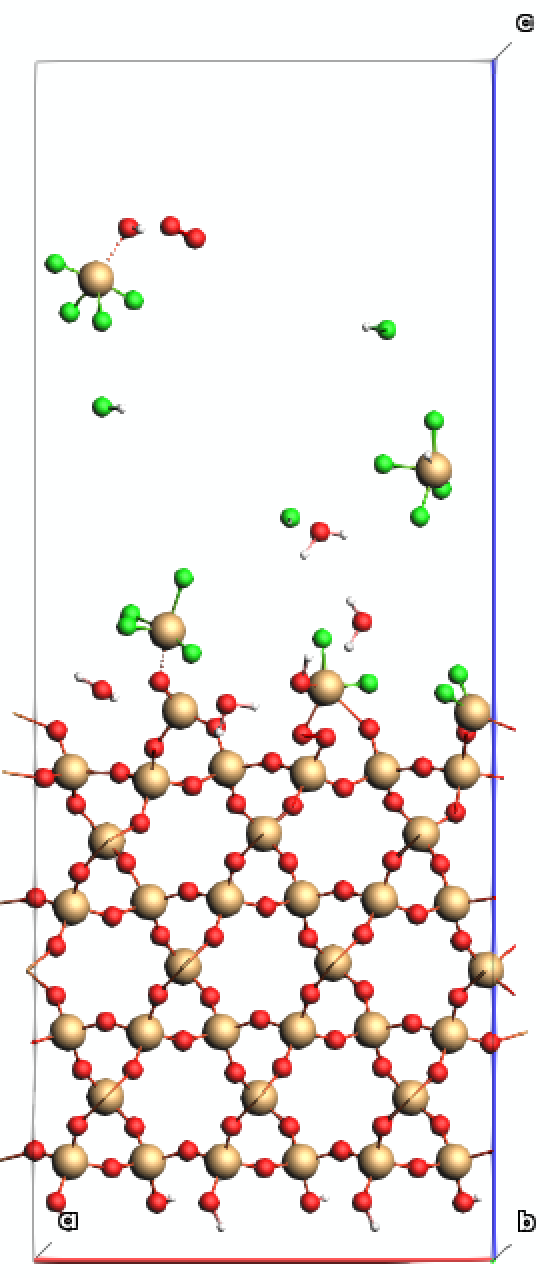
Atomic layer etching (ALE) of SiO2 using HF involves a self-limiting, two-step process to achieve precise, angstrom-level removal. It typically employs sequential exposure to hydrogen fluoride (HF) and a second reactant (like Trimethylaluminum) to modify the surface and remove the modified layer. In this tutorial, the first half-cycle is represented by HF molecules impinging on a hydroxylated SiO2 slab.
The model uses three main ingredients:
a hydroxylated SiO2 slab,
a frozen bottom region that anchors the slab and a thermostatted active region, and
an HF molecule gun that inserts projectiles above the slab with a downward velocity.
The workflow consists of a ReaxFF geometry optimization followed by a ReaxFF molecular dynamics calculation with molecule insertion and removal.
Part A: Prepare the Hydroxylated Slab¶
The notebook constructs the SiO2 slab from lattice vectors, cuts a surface, trims atoms at the top and bottom, and adds hydrogens to hydroxylate both exposed sides. Here we will start from the prepared XYZ file. You can also skip the relaxation and download the relaxed slab at the end of part C below.
Import the prepared slab¶
SiO2_OH_slab_001.xyzThe molecular dynamics calculation treats the slab in two regions:
frozen: atoms near the bottom of the slab, fixed during MD,thermo: all remaining slab atoms, controlled by a thermostat during MD.
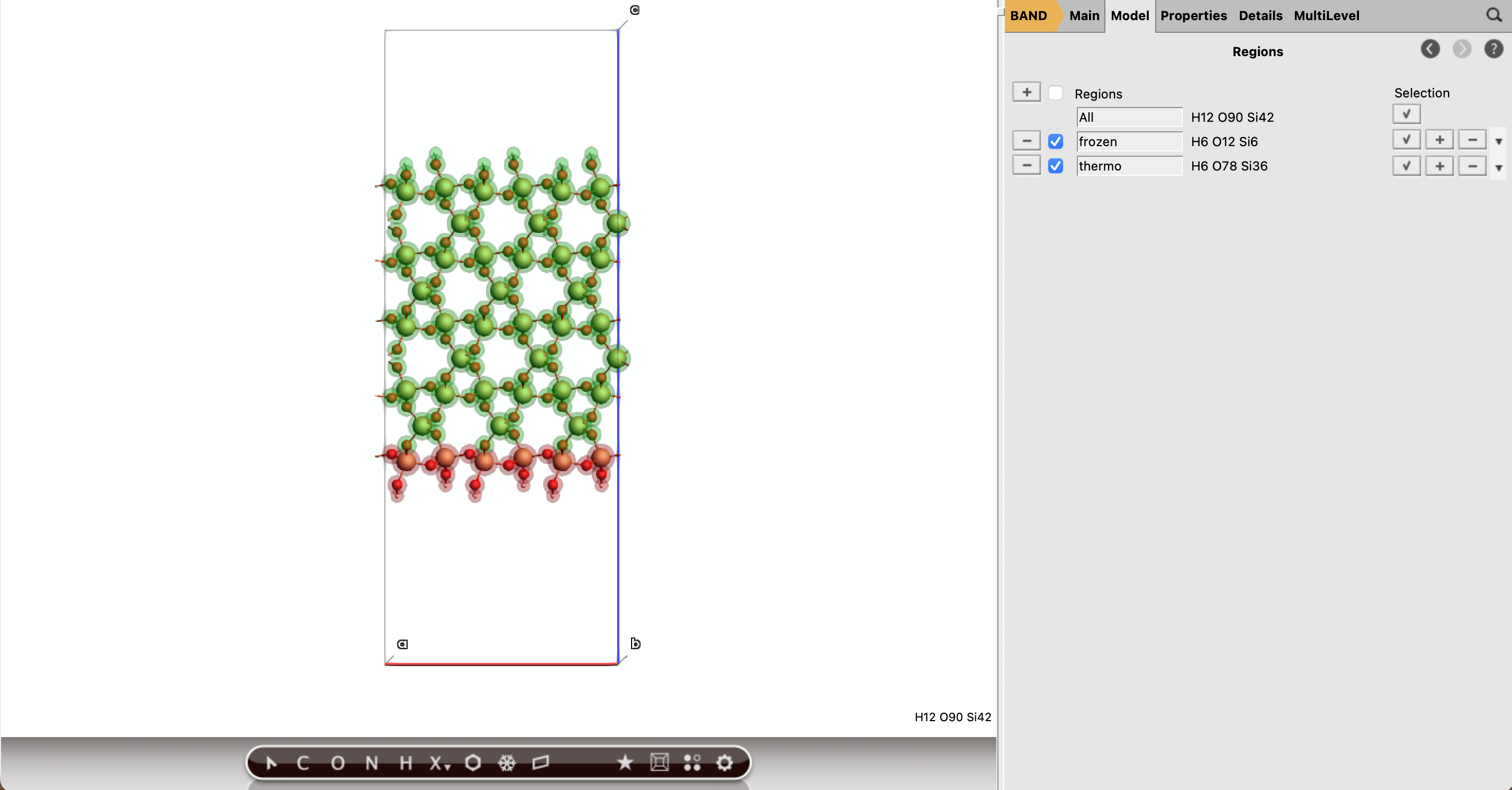
Part A: Assign the regions¶
3 Å of the bottom of the slab (at least include the first bottom layer of Si)frozenfrozen region you can Select → Invert Selection)thermoSiO2_slab_relaxTip
The exact atom selection can be checked visually. The frozen region should form a thin support layer at the bottom of the slab, while the exposed upper surface should belong to thermo.
Part C: Relax the Slab¶
Before shooting HF projectiles at the surface, relax the hydroxylated slab with ReaxFF. This removes artificial strain introduced by slab construction and hydrogen placement.
Set up the ReaxFF geometry optimization¶
 →
→ 
HONSiF.ffAfter the job finishes, open the optimized structure in AMSinput, update the molecule and save it for the MD calculation. The prepared relaxed structure is also available as SiO2_OH_slab_001_relaxed.xyz.
Part D: Create the HF Projectile¶
F key, click above the slab to create a F atom, and press cmd/ctrl+E to passivate F and create HFHF region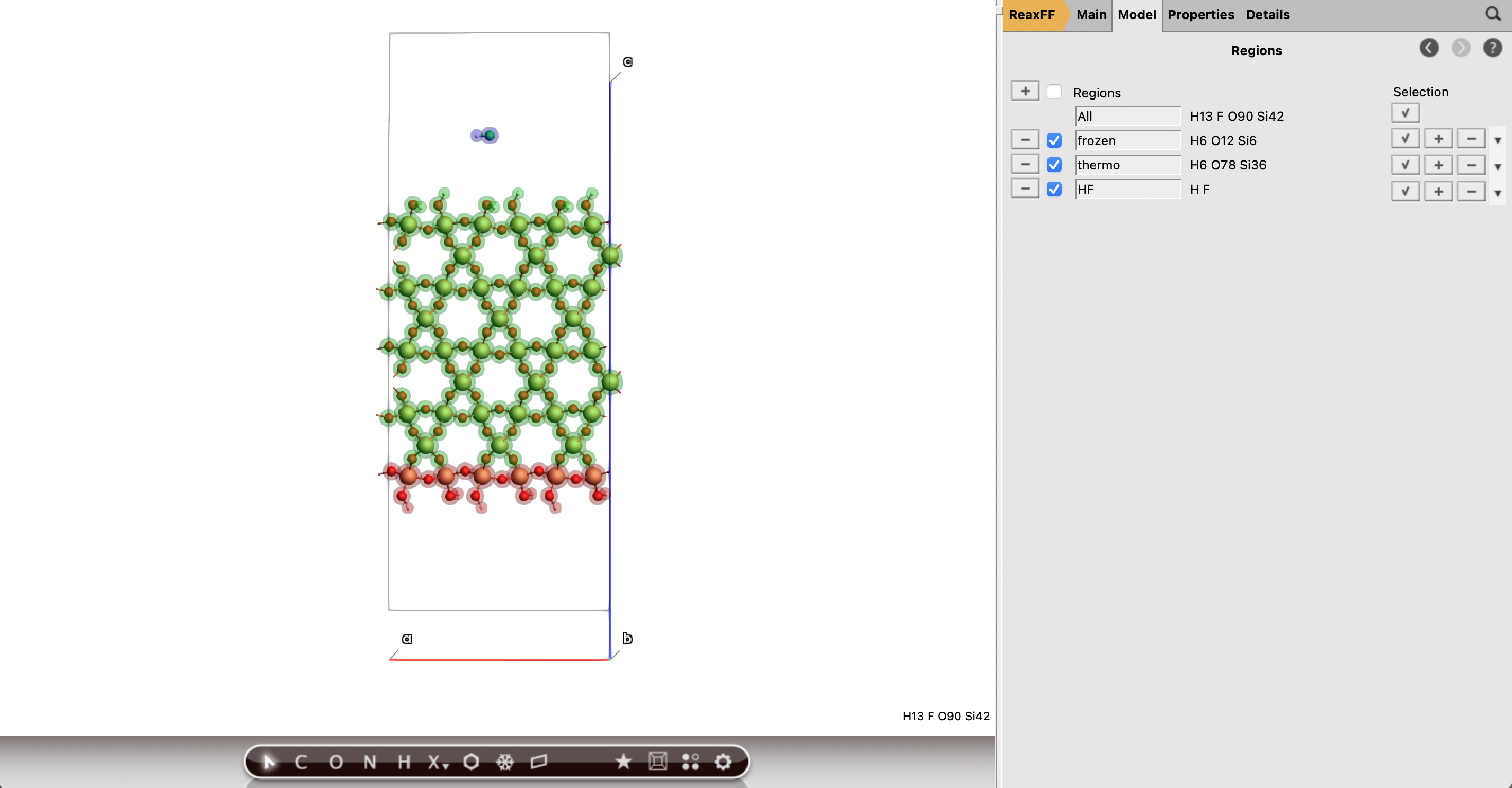
Part E: Configure Molecule-Gun MD¶
The half-cycle simulation inserts HF molecules above the slab and gives them a downward velocity. Molecules that leave the active region are removed with sink boxes.
Set the molecular dynamics task¶
HONSiF.ff force field100000100 K10000, to save 10 checkpoint filesConfigure the thermostat and constraints¶
NHC300 K100 fsthermo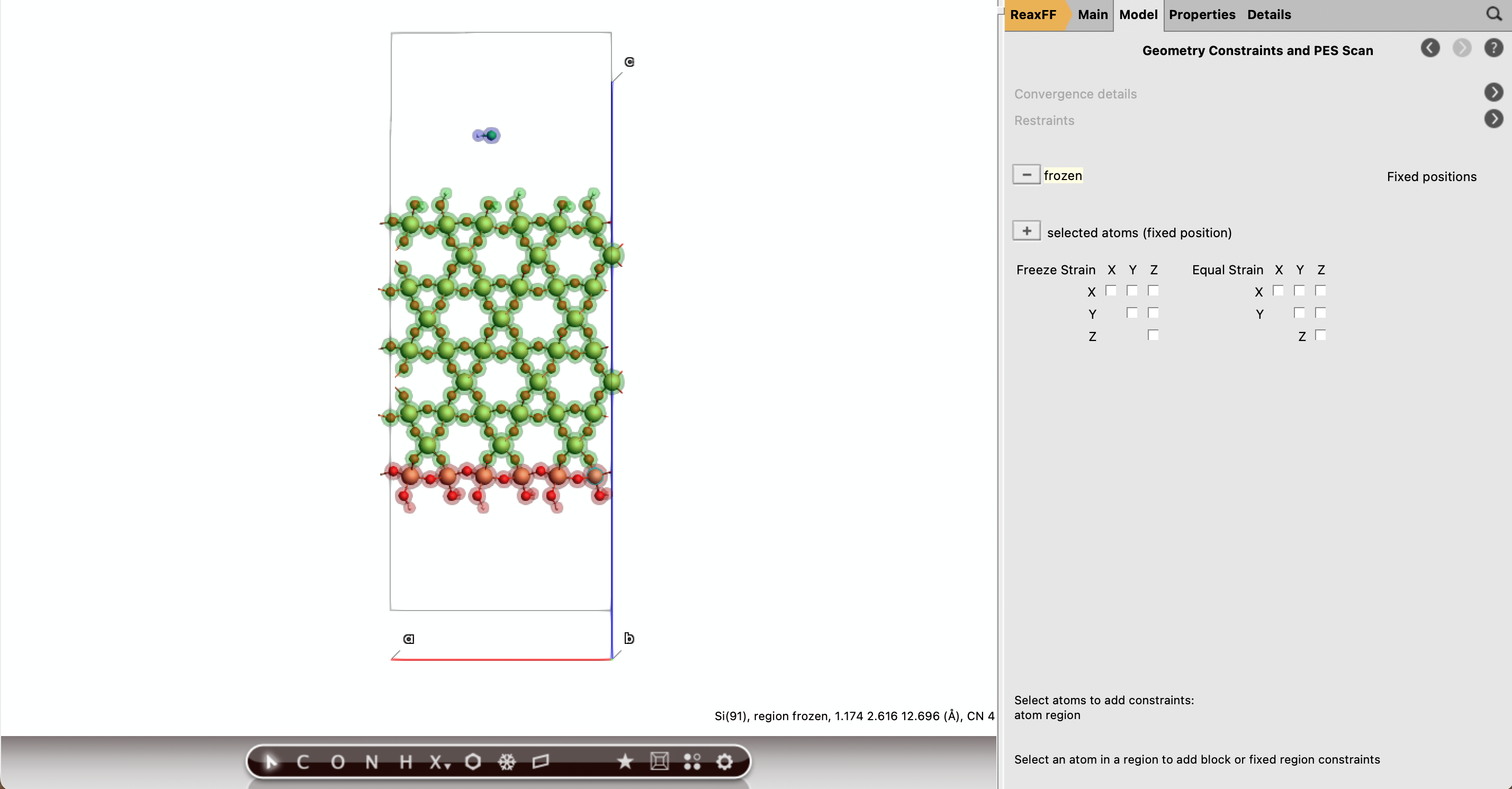
Add the HF molecule gun¶
Select the slab and move it such that the bottom of the slab is slightly above the z = 0 coordinates. Then you can Edit → Crystal → Map Atoms To (0..1) to have a clean setup.
HF10001000300 K3.0 Å0 1 0 1 0.7 0.80.1 Å/fs60 degreesTip
The insertion box should be placed above the relaxed surface, not inside the slab. In the notebook this position is computed from the top of the relaxed slab plus 6 Å.
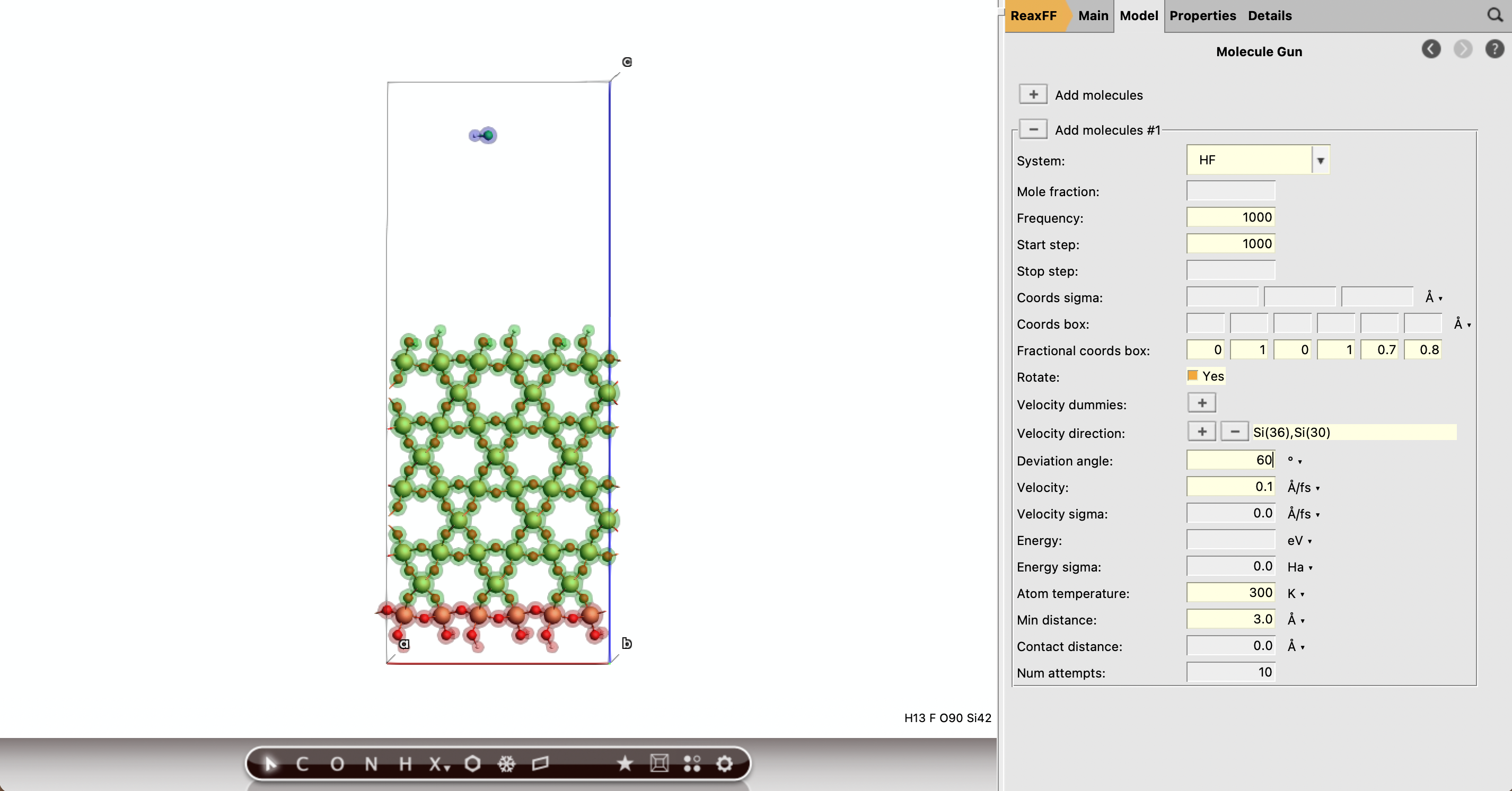
Add the removal sink boxes¶
*10.0 1.0 0.0 1.0 0.95 0.98*10.0 1.0 0.0 1.0 0.05 0.02Note
Be sure that the bottom of the slab is above the fractional coordinate 0.05 or the job might crash trying to remove an atom from a fixed region.
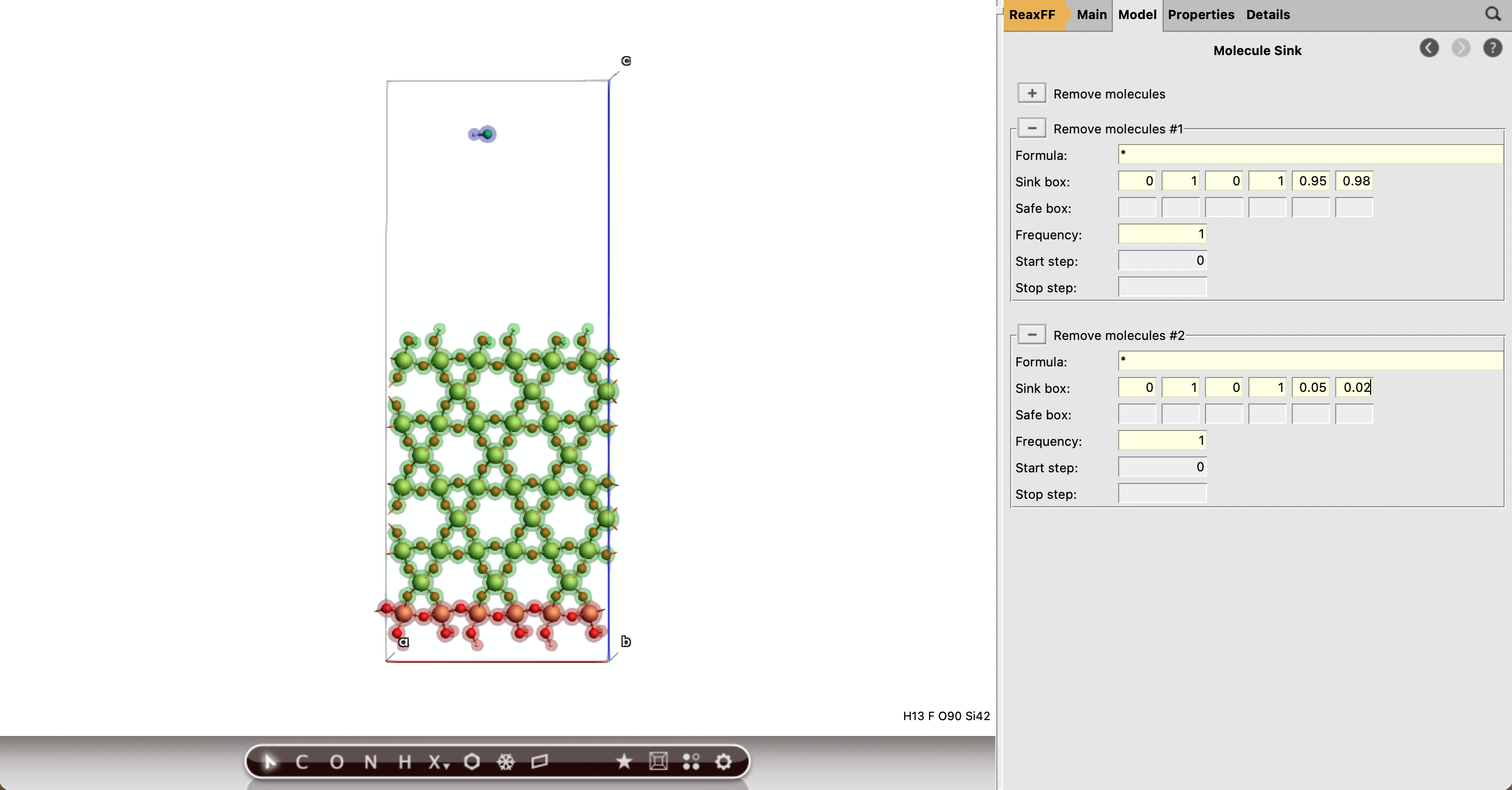
Run the calculation¶
Part F: Inspect the Trajectory¶
After the MD calculation has produced trajectory frames, inspect representative structures in AMSmovie.
Analyze the molecule-gun trajectory¶
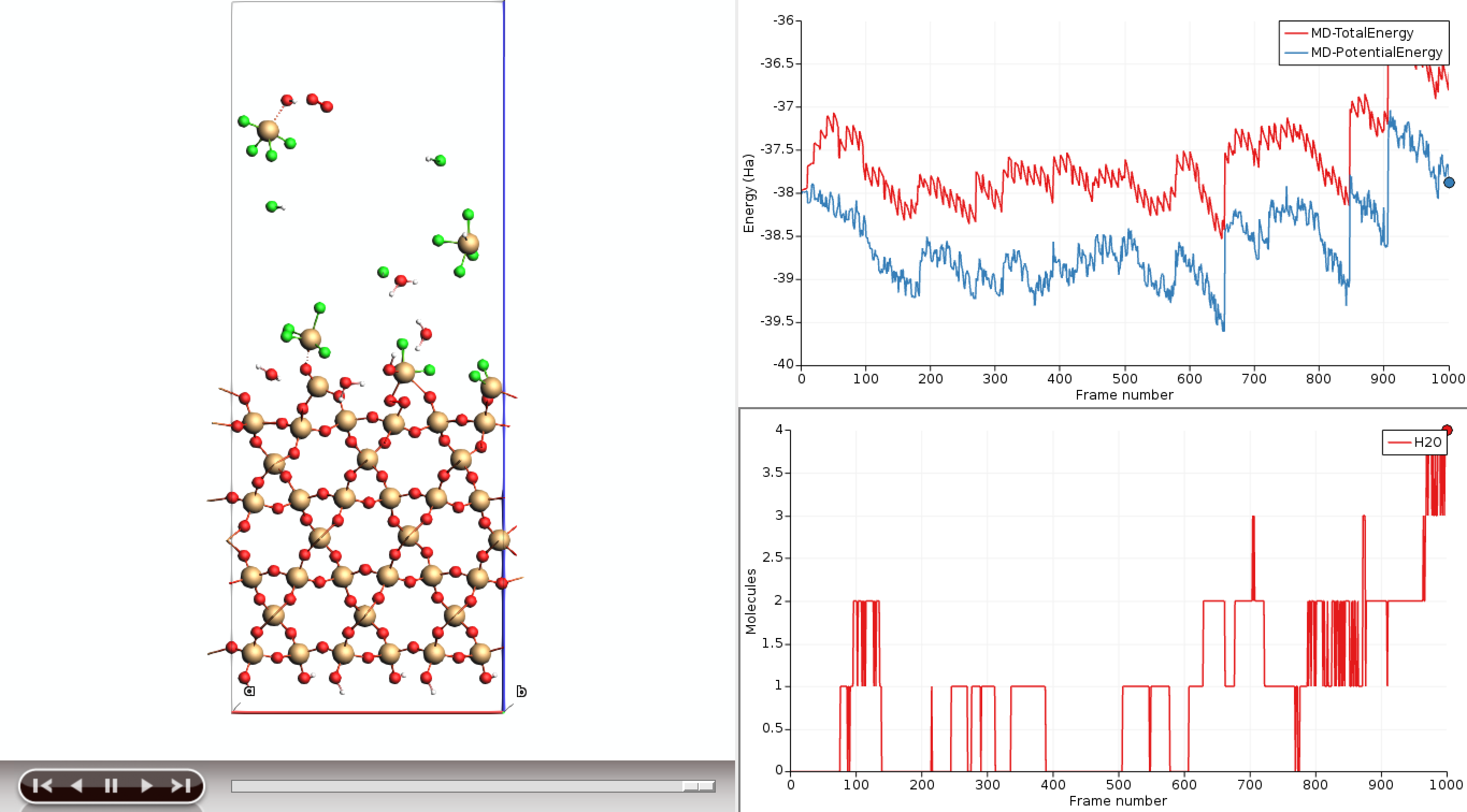
Note
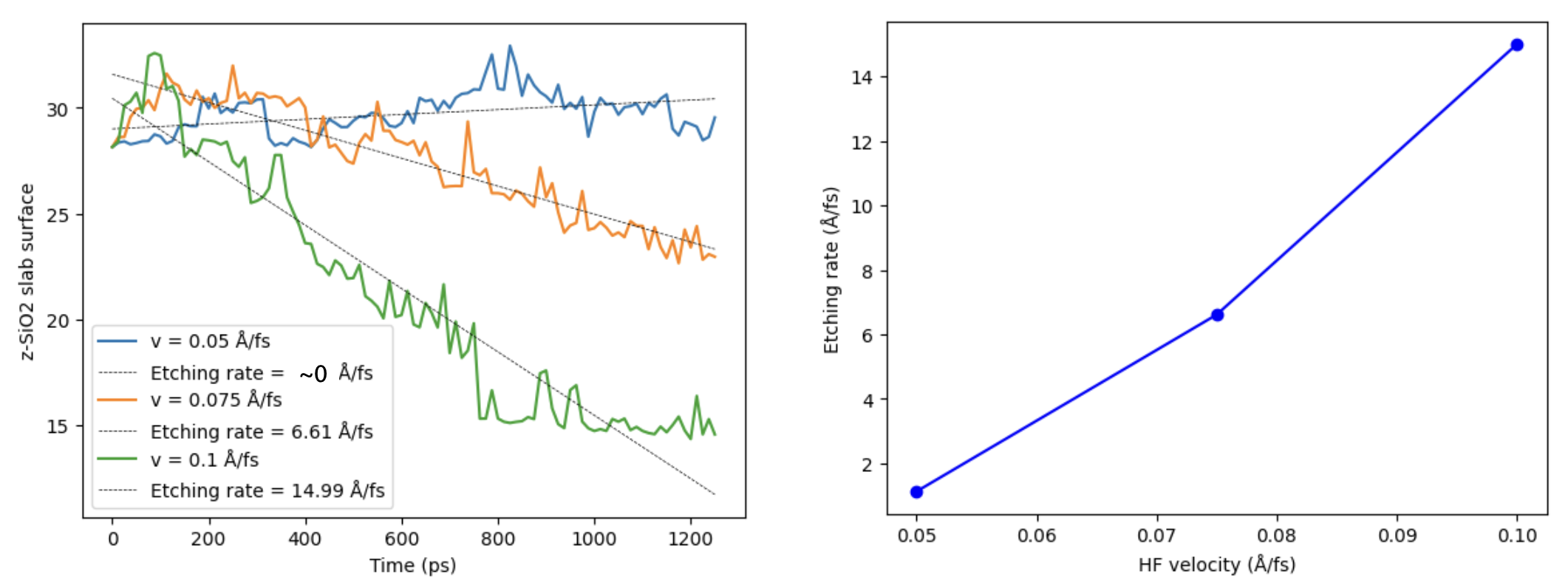
You usually want to run this simulation for many more steps to observe some important etching of the surface. The etching rates below were computed based on simulations of 1 ns.
Summary¶
The same setup can be adapted to other projectile molecules, slab sizes, impact velocities, and surface preparations. You can measure the top of the slab as a function of time and extract etching rates.