Correlation Plots for Computed Results¶
This example illustrates how to plot, for example, forces predicted by one engine against forces predicted by another.
Initial imports¶
import scm.plams as plams
from scm.plams.tools.plot import plot_correlation, get_correlation_xy
import matplotlib.pyplot as plt
Define two engines to compare¶
Here we choose GFNFF and GFN1-xTB
e1 = plams.Settings()
e1.input.GFNFF
e2 = plams.Settings()
e2.input.DFTB.Model = "GFN1-xTB"
Let’s use a glycine molecule generated from SMILES:
from scm.base import ChemicalSystem
glycine = ChemicalSystem.from_smiles("C(C(=O)O)N")
plams.view(glycine, width=200, height=200, padding=-0.5)
Run a single-point calculation storing the Gradients (negative forces):
sp = plams.Settings()
sp.input.ams.Task = "SinglePoint"
sp.input.ams.Properties.Gradients = "Yes"
sp.runscript.nproc = 1 # run in serial
job1 = plams.AMSJob(settings=sp + e1, name="glycine-engine1", molecule=glycine)
job2 = plams.AMSJob(settings=sp + e2, name="glycine-engine2", molecule=glycine)
job1.run()
job2.run()
[18.03|15:53:50] JOB glycine-engine1 STARTED
[18.03|15:53:50] JOB glycine-engine1 RUNNING
[18.03|15:53:50] JOB glycine-engine1 FINISHED
[18.03|15:53:50] JOB glycine-engine1 SUCCESSFUL
[18.03|15:53:50] JOB glycine-engine2 STARTED
[18.03|15:53:50] JOB glycine-engine2 RUNNING
[18.03|15:53:50] JOB glycine-engine2 FINISHED
[18.03|15:53:50] JOB glycine-engine2 SUCCESSFUL
<scm.plams.interfaces.adfsuite.ams.AMSResults at 0x174a7ae20>
plot_correlation(
job1,
job2,
section="AMSResults",
variable="Gradients",
file="engine",
);
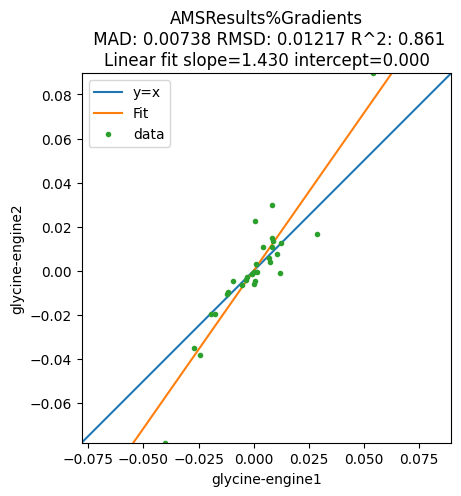
To get the actual numbers, use get_correlation_xy:
x, y = plams.tools.plot.get_correlation_xy(
job1, job2, section="AMSResults", variable="Gradients", file="engine"
)
print("x")
print(x)
print("y")
print(y)
x
[ 1.19424632e-03 1.92134706e-04 6.66726815e-04 -2.68722244e-02
-9.34887191e-04 -4.01123910e-02 1.17988215e-02 4.31998219e-03
5.43487308e-02 8.33720932e-03 5.26401890e-04 1.24250877e-02
8.91893628e-03 8.12876763e-03 2.87360972e-02 -4.14954879e-05
-3.23202276e-03 -9.44698138e-03 -3.49925920e-03 8.54464635e-03
1.69353194e-03 1.06479919e-02 -5.23262379e-03 -2.43272087e-02
7.04191867e-03 -1.95558968e-02 -1.18022032e-02 -1.75261448e-02
7.24349781e-03 -1.21813902e-02]
y
[ 0.00326195 -0.00599574 0.02250686 -0.03506961 -0.00156447 -0.07804685
-0.0008342 0.01094663 0.08968399 0.02986723 -0.00458719 0.01262981
0.01335181 0.01483532 0.01659624 -0.00041231 -0.0029202 -0.00449173
-0.00404233 0.01096039 -0.00074098 0.00750758 -0.006259 -0.03825885
0.00589418 -0.01943988 -0.00943605 -0.01952429 0.00402414 -0.01044246]
Compare multiple jobs¶
smiles_list = ["CC=C", "CCCO", "C(C(=O)O)N"]
names = ["propene", "propanol", "glycine"]
molecules = [ChemicalSystem.from_smiles(x) for x in smiles_list]
imgs = {n: plams.view(mol, width=200, height=200) for n, mol in zip(names, molecules)}
plams.plot_image_grid(imgs, rows=1);
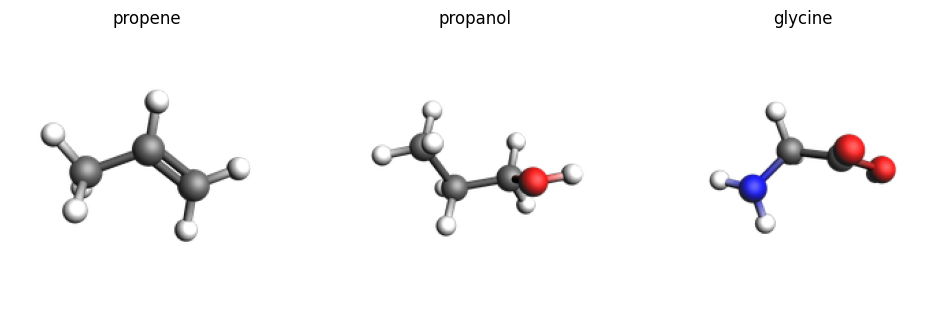
jobs1 = [
plams.AMSJob(settings=sp + e1, name="e1" + name, molecule=mol)
for name, mol in zip(names, molecules)
]
jobs2 = [
plams.AMSJob(settings=sp + e2, name="e2" + name, molecule=mol)
for name, mol in zip(names, molecules)
]
for job in jobs1 + jobs2:
job.run()
[18.03|15:53:53] JOB e1propene STARTED
[18.03|15:53:53] JOB e1propene RUNNING
[18.03|15:53:53] JOB e1propene FINISHED
[18.03|15:53:53] JOB e1propene SUCCESSFUL
[18.03|15:53:53] JOB e1propanol STARTED
[18.03|15:53:53] JOB e1propanol RUNNING
[18.03|15:53:53] JOB e1propanol FINISHED
[18.03|15:53:53] JOB e1propanol SUCCESSFUL
[18.03|15:53:53] JOB e1glycine STARTED
[18.03|15:53:53] Job e1glycine previously run as glycine-engine1, using old results
... output trimmed ....
[18.03|15:53:53] JOB e2propene RUNNING
[18.03|15:53:54] JOB e2propene FINISHED
[18.03|15:53:54] JOB e2propene SUCCESSFUL
[18.03|15:53:54] JOB e2propanol STARTED
[18.03|15:53:54] JOB e2propanol RUNNING
[18.03|15:53:54] JOB e2propanol FINISHED
[18.03|15:53:54] JOB e2propanol SUCCESSFUL
[18.03|15:53:54] JOB e2glycine STARTED
[18.03|15:53:54] Job e2glycine previously run as glycine-engine2, using old results
[18.03|15:53:54] JOB e2glycine COPIED
The correlation plot can be plotted as before. You can also add a unit conversion to get your preferred units, and add custom xlabel and ylabel:
unit = "eV/angstrom"
multiplier = plams.Units.convert(1.0, "hartree/bohr", unit)
plot_correlation(
jobs1,
jobs2,
section="AMSResults",
variable="Gradients",
file="engine",
xlabel="Engine 1",
ylabel="Engine 2",
unit=unit,
multiplier=multiplier,
);
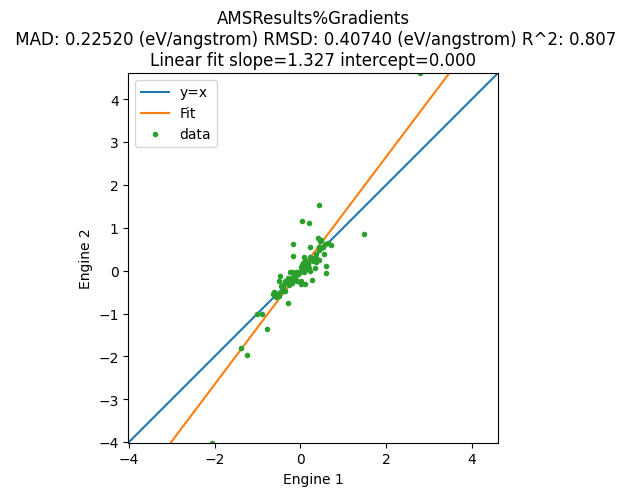
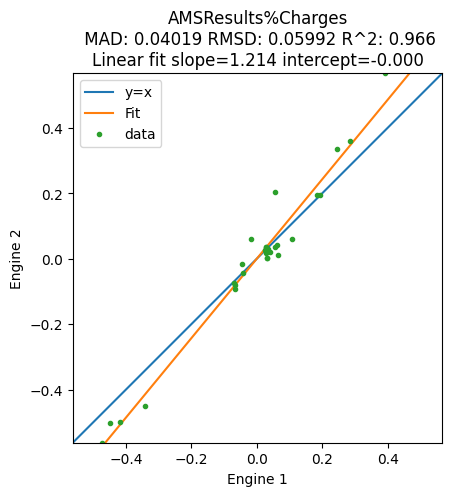
plot_correlation(
jobs1,
jobs2,
section="AMSResults",
variable="Charges",
file="engine",
xlabel="Engine 1",
ylabel="Engine 2",
);
Use Task Replay to compare multiple frames from a trajectory¶
The forces from an MD job can be stored with writeenginegradients=True
md = plams.AMSNVEJob(
settings=e1,
name="nve-md-e1",
molecule=glycine,
velocities=400,
nsteps=100,
samplingfreq=10,
writeenginegradients=True,
)
md.run();
[18.03|15:53:54] JOB nve-md-e1 STARTED
[18.03|15:53:54] JOB nve-md-e1 RUNNING
[18.03|15:53:55] JOB nve-md-e1 FINISHED
[18.03|15:53:55] JOB nve-md-e1 SUCCESSFUL
When using the Replay task, set Properties.Gradients to get the forces:
replay_s = plams.Settings()
replay_s.input.ams.Task = "Replay"
replay_s.input.ams.Properties.Gradients = "Yes"
replay_s.input.ams.Replay.File = md.results.rkfpath()
replay = plams.AMSJob(settings=e2 + replay_s, name="replay-e2")
replay.run();
[18.03|15:53:55] JOB replay-e2 STARTED
[18.03|15:53:55] JOB replay-e2 RUNNING
[18.03|15:53:56] JOB replay-e2 FINISHED
[18.03|15:53:56] JOB replay-e2 SUCCESSFUL
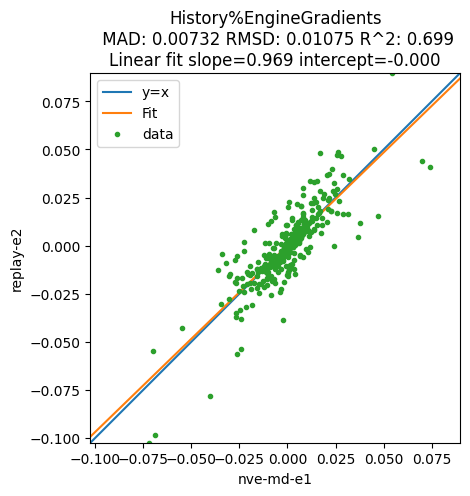
For the MD job the gradients (negative forces) are stored in History%EngineGradients, but for the Replay job they are stored in History%Gradients. Use the alt_variable to specify the variable for the second job:
plot_correlation(
md, replay, section="History", variable="EngineGradients", alt_variable="Gradients", file="ams"
);
See also¶
Python Script¶
#!/usr/bin/env python
# coding: utf-8
# ## Initial imports
import scm.plams as plams
from scm.plams.tools.plot import plot_correlation, get_correlation_xy
import matplotlib.pyplot as plt
# ## Define two engines to compare
#
# Here we choose GFNFF and GFN1-xTB
e1 = plams.Settings()
e1.input.GFNFF
e2 = plams.Settings()
e2.input.DFTB.Model = "GFN1-xTB"
# Let's use a glycine molecule generated from SMILES:
from scm.base import ChemicalSystem
glycine = ChemicalSystem.from_smiles("C(C(=O)O)N")
plams.view(glycine, width=200, height=200, padding=-0.5)
# Run a single-point calculation storing the Gradients (negative forces):
sp = plams.Settings()
sp.input.ams.Task = "SinglePoint"
sp.input.ams.Properties.Gradients = "Yes"
sp.runscript.nproc = 1 # run in serial
job1 = plams.AMSJob(settings=sp + e1, name="glycine-engine1", molecule=glycine)
job2 = plams.AMSJob(settings=sp + e2, name="glycine-engine2", molecule=glycine)
job1.run()
job2.run()
plot_correlation(
job1,
job2,
section="AMSResults",
variable="Gradients",
file="engine",
)
# To get the actual numbers, use ``get_correlation_xy``:
x, y = plams.tools.plot.get_correlation_xy(job1, job2, section="AMSResults", variable="Gradients", file="engine")
print("x")
print(x)
print("y")
print(y)
# ## Compare multiple jobs
smiles_list = ["CC=C", "CCCO", "C(C(=O)O)N"]
names = ["propene", "propanol", "glycine"]
molecules = [ChemicalSystem.from_smiles(x) for x in smiles_list]
imgs = {n: plams.view(mol, width=200, height=200) for n, mol in zip(names, molecules)}
plams.plot_image_grid(imgs, rows=1)
jobs1 = [plams.AMSJob(settings=sp + e1, name="e1" + name, molecule=mol) for name, mol in zip(names, molecules)]
jobs2 = [plams.AMSJob(settings=sp + e2, name="e2" + name, molecule=mol) for name, mol in zip(names, molecules)]
for job in jobs1 + jobs2:
job.run()
# The correlation plot can be plotted as before. You can also add a unit conversion to get your preferred units, and add custom xlabel and ylabel:
unit = "eV/angstrom"
multiplier = plams.Units.convert(1.0, "hartree/bohr", unit)
plot_correlation(
jobs1,
jobs2,
section="AMSResults",
variable="Gradients",
file="engine",
xlabel="Engine 1",
ylabel="Engine 2",
unit=unit,
multiplier=multiplier,
)
plot_correlation(
jobs1,
jobs2,
section="AMSResults",
variable="Charges",
file="engine",
xlabel="Engine 1",
ylabel="Engine 2",
)
# ## Use Task Replay to compare multiple frames from a trajectory
# The forces from an MD job can be stored with ``writeenginegradients=True``
md = plams.AMSNVEJob(
settings=e1,
name="nve-md-e1",
molecule=glycine,
velocities=400,
nsteps=100,
samplingfreq=10,
writeenginegradients=True,
)
md.run()
# When using the Replay task, set ``Properties.Gradients`` to get the forces:
replay_s = plams.Settings()
replay_s.input.ams.Task = "Replay"
replay_s.input.ams.Properties.Gradients = "Yes"
replay_s.input.ams.Replay.File = md.results.rkfpath()
replay = plams.AMSJob(settings=e2 + replay_s, name="replay-e2")
replay.run()
# For the MD job the gradients (negative forces) are stored in ``History%EngineGradients``, but for the Replay job they are stored in ``History%Gradients``. Use the ``alt_variable`` to specify the variable for the second job:
plot_correlation(md, replay, section="History", variable="EngineGradients", alt_variable="Gradients", file="ams")