3. Python Library¶
Setting up and running ACErxn with the ACErxn Python library consists of five steps.
Creating reactant and product PLAMS
Moleculeobjects.Creating a
Reactantobject from the list of reactantMoleculeobjects, their charges, and a list of selected active atoms.Creating a PLAMS
Settingsobject containing any non-default ACErxn settings.Creating an
ACErxnJobobject from the combinedReactantobject, productMoleculeobject, andSettingsobject.Running the
ACErxnJobobject.
Each of these steps will be addressed in more detail in the following subsections.
3.1. Molecule information¶
The most important part of the input is the molecular information about the reactants and the products.
They need to be provided as PLAMS Molecule objects, which can be read from many formats (e.g. .xyz, .mol).
If the input Molecule objects contain no bonding, then the bonding will be guessed by the ACErxn code (guess_bonds()).
from scm.plams import Molecule
reactant_filenames = ['reactant.xyz']
product_filenames = ['product.xyz']
reactants = [Molecule(fn) for fn in reactant_filenames]
products = [Molecule(fn) for fn in product_filenames]
3.1.1. Reactant Object¶
The ACErxn process requires a lot of additional information on the molecular systems:
What the smallest possible fragments of the system are. The bonds between the atoms in these fragments will never be broken.
The charges of these smallest possible fragments.
The stability of these smallest possible fragments. If a fragment is labeled unstable, it will be required to undergo change in every elementary reaction of the network.
The above information can be guessed automatically by the Reactant object with only minimal user-provided data.
To set up a Reactant object, not only the reactant Molecule objects and their charges are required,
but also a list of active atoms for each reactant molecule.
An active atom is an atom that can partake in bond breaking/forming.
from scm.acerxn import Reactant
molcharges = [0]
active_atoms = [[0,12,13,15]]
# Create the reactant object
reactant = Reactant(reactants, active_atoms, molcharges)
reactant.guess_fragments()
In some cases, two atoms need to be considered active, even though the bond between them should never break. An example would be a dihydroxy-alkane, where the C-OH bonds can be broken, and therefore all C and O atoms must be active. However, the C-C bond is expected to be stable. In that case, the carbon-carbon bond can be specified as frozen in the system block.
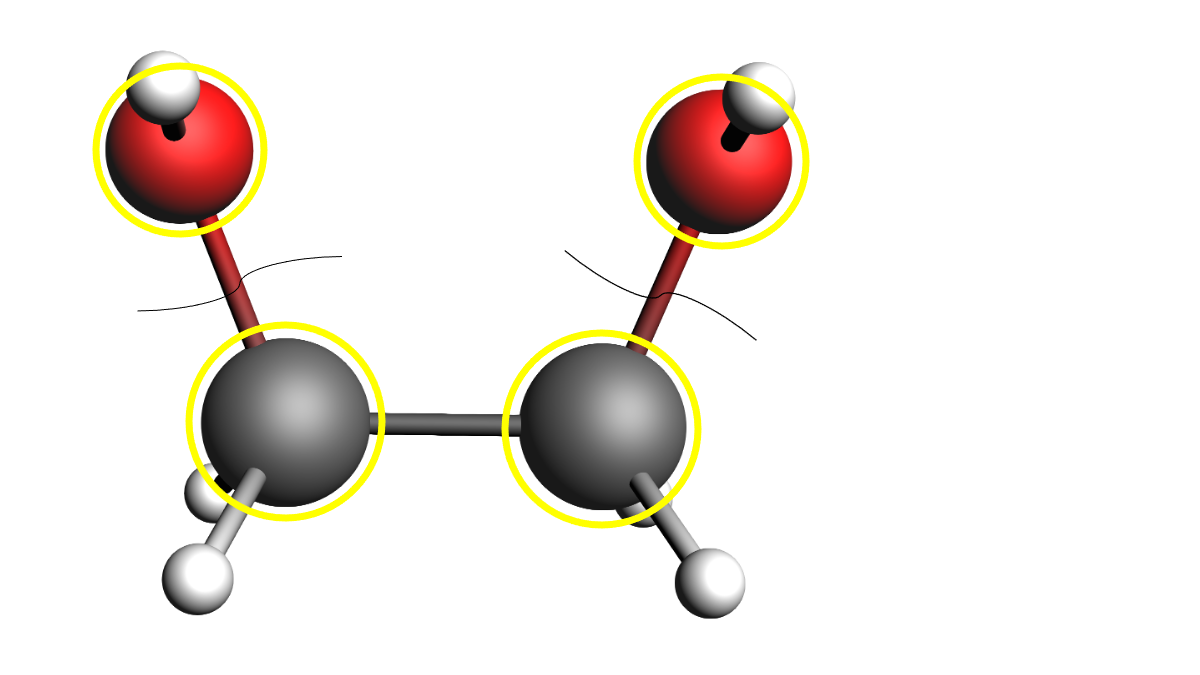
# Read the reactant molecules, and set up their info
reactants = [Molecule(fn) for fn in ['dihydroxyethylene']]
molcharges = [0]
active_atoms = [[0,1,2,3]]
# Create the reactant object
reactant = Reactant(reactants, active_atoms, molcharges)
reactant.guess_fragments(frozen_bonds=[[(0,1)]])
After guess_fragments() is called, all fragment information is created. You can then check that information and alter it if needed.
# Write the fragments as XYZ file and print their charges and stabilities
for i,fragment in enumerate(reactant.fragments) :
fragment.write('%s.xyz'%(fragment.properties.name))
print ('Charge : ',i,fragment.properties.charge)
print ('Stable : ',i,fragment.properties.stable)
3.1.2. Other Molecule objects¶
Apart from the mandatory product molecules, the user can provide intermediates.
ACErxn will then attempt to find reaction paths that include all these intermediates.
We define a single intermediate as a stable (set of) molecule(s) with the same stoichiometry as the combined reactant molecules.
The user can provide many intermediates, and define a Molecule object for each intermediate submolecule.
known_intermediate_filenames = [['IM0_0.xyz', 'IM0_1.xyz', ],['IM1_0.xyz']]
intermediates = [[Molecule(fn) for fn in filelist] for filelist in known_intermediate_filenames]
It is also possible to provide known single molecules and forbidden molecules that do not combine to have the same stoichiometry as the combined reactant molecules.
known_molecules = [Molecule(fn) for fn in ['mol1.xyz','mol2.xyz']]
forbidden_molecules = [Molecule(fn) for fn in ['mol3.xyz','mol4.xyz']]
These can later be passed to the ACErxnJob object.
3.2. Settings ACErxn¶
The settings need to be provided as a PLAMS Settings object. The keywords are described in a later section,
with only a short example shown here.
from scm.plams import Settings
settings = Settings()
settings.RunInfo.Steps = "GenerateIntermediates"
3.3. Settings AMS¶
For the QM geometry optimizations in the first step of ACErxn (intermediate generation),
AMS engine information needs to be added to the Settings object.
Optionally, an additional ForceField engine can be provided, which
will then be used for the initial geometry optimization.
If not provided, UFF with default settings will be used.
Currently UFF is the only engine that can be selected
for the initial optimization.
settings.qmengine.Mopac.Model = 'PM6'
settings.mmengine.ForceField.Type = 'UFF'
settings.mmengine.ForceField.UFF.Library = 'UFF4MOF'
3.4. Setting up and Running the Job¶
Finally, all the input information above can be combined in an ACErxnJob object, which behaves like a PLAMS Job object and can then be directly run. As a result, the reaction network is written to RKF files and can be viewed with AMSMovie or accessed via the result object.
from scm.plams import init, finish
from scm.acerxn import ACErxnJob
# Set up the ACE job
job = ACErxnJob(reactant, products, intermediates, settings=settings)
# Now run the job, and create the reaction network
init()
result = job.run()
finish()
If you want to add known_molecules and forbidden_molecules, they can be added as object instances of ACErxnJob before run() is called.
job.known_molecules = known_molecules
job.forbidden_molecules = forbidden_molecules
The following minimal Python script can be run with amspython.
import networkx as nx
import matplotlib.pyplot as plt
from scm.plams import Molecule, Settings
from scm.plams import init, finish
from scm.acerxn import Reactant, ACErxnNetwork
def main():
"""
The main script
"""
reactant_filenames = ["reactant.xyz"]
product_filenames = ["product.xyz"]
# Some user supplied info on the reactants
molcharges = [0]
active_atoms = [[0, 12, 13, 15]]
# Create the reactant molecules
reactants = [Molecule(fn) for fn in reactant_filenames]
reactant = Reactant(reactants, active_atoms, molcharges)
reactant.guess_fragments()
# Create the product molecules
products = [Molecule(fn) for fn in product_filenames]
# Create the settings object
settings = ACErxnNetwork.default_settings()
# QM engine settings
engine_settings = Settings()
engine_settings.Mopac.Model = "PM6"
settings.qmengine = engine_settings
# Set up the ACE network
network = ACErxnNetwork(reactant, products, settings=settings)
# Now run the network, and create the reaction network
init()
network.generate()
finish()
# Optionally plot the resulting reaction network
# nx.draw( network.graph, with_labels=True, node_color='white', edgecolors='blue', font_weight='bold', node_size=4000 )
# plt.show()
if __name__ == "__main__":
main()
3.5. Components¶
The Reactant and ACErxnJob are the only ACErxn classes needed to create an ACErxn network, and will be discussed in detail first.
After running the job, all network information is stored in a results folder, where it can be accessed by the user, and studied.
Alternatively, it is also possible to explore the resulting networks using Python scripting, using the ACErxnResults and the ACErxnNetwork classes, which are also discussed in the following sections.
Finally, a few available functions are discussed that allow interaction with the specific Settings object used in ACErxn.