Active Learning: Ru/H Part 2, Surface PES Scans¶
Continue the Ru/H case study by constructing Ru slabs, adding adsorbates, and extending the reference data with surface coordinate and bond scans. Run part 1 first.
Downloads: Notebook | Script ?
Additional files required by this example:
common_ru_h.py
Requires: AMS2026 or later
Required AMS Packages: m3gnet
Related examples
Related tutorials
Related documentation

Initial imports¶
import scm.plams as plams
from scm.params import ResultsImporter
from scm.plams import Settings, AMSJob, log, Molecule
from pathlib import Path
import matplotlib.pyplot as plt
# common_ru_h.py must exist in the current working directory
from common_ru_h import rotation, check_installation
# register dependencies for AMSjobs, to support submitting this notebook directly to a cluster in AMS2025+
# dependency: {} common_ru_h.py
# dependency: {} reference_data_1
Initialize PLAMS working directory¶
old_ref_dir = "reference_data_1"
check_installation(old_ref_dir)
new_ref_dir = "reference_data_2"
ri = ResultsImporter.from_yaml(old_ref_dir)
plams.init()
Current AMS version: 2024.102
05-31 11:15:26 m3gnet is installed: M3GNet ML Backend v[0.2.4] - build:0 [06668e0a45ce742d8f66ff23484b8a1e]
05-31 11:15:26 qe is installed: Quantum ESPRESSO (AMSPIPE) v[7.1] - build:115 [777d72eb480fe4d632a003cc62e9c1cb]
PLAMS working folder: /path/to/plams_workdir.002
Load the optimized bulk Ru structure from the job collection¶
The lattice was optimized in the previous notebook, and the structure was stored in the job collection.
Let’s retrieve it from the job collection and use it to construct Ru surface slabs.
optimized_bulk = ri.job_collection["hcp_lattopt_Ru_dft"].molecule
plams.plot_molecule(optimized_bulk, rotation=rotation)
plt.title("Bulk hcp Ru");

Cut out Ru(0001) and Ru(10-10) slabs¶
The PLAMS Molecule class does not have any methods for creating slabs. However, we can use the ChemicalSystem in AMS2024+. The details can be found in the slice_slab() function in common_ru_h.py.
from common_ru_h import slice_slab
slab_100 = slice_slab(
optimized_bulk,
miller=(1, 0, 0),
thickness=7.0,
cell_z=15,
ref_atom=0,
)
slab_100 = slab_100.supercell(3, 2, 1)
for at in slab_100:
at.properties = Settings() # remove details about supercell generation
plams.plot_molecule(slab_100, rotation=rotation)
plt.title("Ru(10-10)");

from common_ru_h import slice_slab
slab_001 = slice_slab(
optimized_bulk,
miller=(0, 0, 1),
thickness=7.0,
cell_z=15,
ref_atom=0,
)
slab_001 = slab_001.supercell([[3, 0, 0], [2, 4, 0], [0, 0, 1]])
for at in slab_001:
at.properties = Settings() # remove details about supercell generation
plams.plot_molecule(slab_001, rotation=rotation)
plt.title("Ru(0001)");

This slab is perhaps too thick, so let’s remove the top atomic layer:
for at in slab_001:
if at.coords[2] > 10:
slab_001.delete_atom(at)
plams.plot_molecule(slab_001, rotation=rotation)
plt.title("Ru(0001)");

Add hydrogens¶
from common_ru_h import add_adsorbate
slab_100_H = add_adsorbate(slab_100, "H", frac_x=0, frac_y=0, delta_z=2.0)
# add second adsorbate, now delta_z=2.0 means 2 angstrom above the previous H
slab_100_H2 = add_adsorbate(slab_100_H, "H", frac_x=0.5, frac_y=0.5, delta_z=2.0)
plams.plot_molecule(slab_100_H2, rotation=rotation, radii=0.6)
plt.title("Ru(10-10) with H");

from common_ru_h import add_adsorbate
slab_001_H = add_adsorbate(slab_001, "H", frac_x=0, frac_y=0, delta_z=2.0)
slab_001_H2 = add_adsorbate(slab_001_H, "H", frac_x=0.5, frac_y=0.5, delta_z=2.0)
plams.plot_molecule(slab_001_H2, rotation=rotation, radii=0.6)
plt.title("Ru(0001) with H");

Set up and run the jobs¶
Ru-H distance bond scan on Ru(10-10)¶
from common_ru_h import get_surface_bond_scan_coordinates
from common_ru_h import (
get_bottom_atom_index,
pesscan_settings,
m3gnet_up_settings,
constraints_settings,
plot_pesscan,
)
system = slab_100_H
scan_coordinates = get_surface_bond_scan_coordinates(
system, atom_index=len(system), start=1.2, end=2.5
)
pesscan_Ru_H_distance = plams.AMSJob(
settings=(
pesscan_settings(scan_coordinates, n_points=6)
+ m3gnet_up_settings()
+ constraints_settings(get_bottom_atom_index(system))
),
name="PESScan_Ru_H_distance_m3gnet",
molecule=system,
)
pesscan_Ru_H_distance.run()
# !amsmovie {pesscan_Ru_H_distance.results.rkfpath()} # open in AMSmovie
plot_pesscan(pesscan_Ru_H_distance);
[31.05|11:15:31] JOB PESScan_Ru_H_distance_m3gnet STARTED
[31.05|11:15:31] JOB PESScan_Ru_H_distance_m3gnet RUNNING
[31.05|11:16:35] JOB PESScan_Ru_H_distance_m3gnet FINISHED
[31.05|11:16:36] Job PESScan_Ru_H_distance_m3gnet reported warnings. Please check the output
[31.05|11:16:36] JOB PESScan_Ru_H_distance_m3gnet SUCCESSFUL

Surface and bulk diffusion H on/in Ru(10-10)¶
Note: These PES scans are just meant to sample H atoms diffusing through the slab and on the surface.
You can also set up more intelligent PES scans (for example, to sample diffusion along specific pathways like hcp-bridge-fcc etc.). In these examples, the H atom simply diffuses from one corner of the cell to the opposite corner.
from common_ru_h import (
get_surface_diffusion_scan_coordinates,
get_bulk_diffusion_scan_coordinates,
)
from common_ru_h import (
get_bottom_atom_index,
pesscan_settings,
m3gnet_up_settings,
constraints_settings,
plot_pesscan,
)
system = slab_100_H2
scan_coordinates = get_surface_diffusion_scan_coordinates(
system, atom_index=len(system) - 1
)
scan_coordinates += get_bulk_diffusion_scan_coordinates(
system, atom_index=len(system), delta_z=2.0
)
pesscan_100_H2 = plams.AMSJob(
settings=(
pesscan_settings(scan_coordinates, n_points=20)
+ m3gnet_up_settings()
+ constraints_settings(get_bottom_atom_index(system))
),
name="PESScan_100_H2_m3gnet",
molecule=system,
)
pesscan_100_H2.run()
# !amsmovie {pesscan_100_H2.results.rkfpath()} # open in AMSmovie
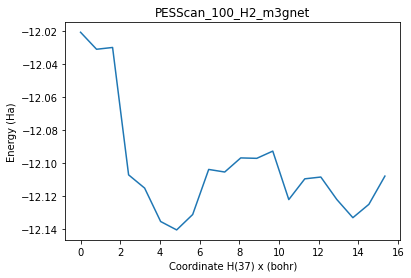
plot_pesscan(pesscan_100_H2);
Atom 37 will diffuse from (x, y) = 0.0, 0.0 to 8.1145, 8.547 with the z coordinate optimized
Atom 38 will diffuse from z = 14.1507 to 2.6849 with the x and y coordinates optimized
[31.05|11:16:36] JOB PESScan_100_H2_m3gnet STARTED
[31.05|11:16:36] JOB PESScan_100_H2_m3gnet RUNNING
[31.05|11:18:14] JOB PESScan_100_H2_m3gnet FINISHED
[31.05|11:18:14] JOB PESScan_100_H2_m3gnet SUCCESSFUL
Surface and bulk diffusion H on/in Ru(0001)¶
from common_ru_h import (
get_surface_diffusion_scan_coordinates,
get_bulk_diffusion_scan_coordinates,
)
from common_ru_h import (
get_bottom_atom_index,
pesscan_settings,
m3gnet_up_settings,
constraints_settings,
plot_pesscan,
)
system = slab_001_H2
scan_coordinates = get_surface_diffusion_scan_coordinates(
system, atom_index=len(system) - 1
)
scan_coordinates += get_bulk_diffusion_scan_coordinates(
system, atom_index=len(system), delta_z=2.0
)
pesscan_001_H2 = plams.AMSJob(
settings=(
pesscan_settings(scan_coordinates, n_points=20)
+ m3gnet_up_settings()
+ constraints_settings(get_bottom_atom_index(system))
),
name="PESScan_001_H2_m3gnet",
molecule=system,
)
pesscan_001_H2.run()
# !amsmovie {pesscan_001_H2.results.rkfpath()} # open in AMSmovie
plot_pesscan(pesscan_001_H2);
Atom 37 will diffuse from (x, y) = 0.0, 0.0 to 8.1145, 9.3698 with the z coordinate optimized
Atom 38 will diffuse from z = 12.5470 to 2.2735 with the x and y coordinates optimized
[31.05|11:18:15] JOB PESScan_001_H2_m3gnet STARTED
[31.05|11:18:15] JOB PESScan_001_H2_m3gnet RUNNING
[31.05|11:19:49] JOB PESScan_001_H2_m3gnet FINISHED
[31.05|11:19:49] JOB PESScan_001_H2_m3gnet SUCCESSFUL

Run DFT singlepoints on PES Scan structures and store on disk¶
from common_ru_h import dft_settings, replay_settings, QEKPointsConfig
replay_jobs = dict()
for pesscan_job in [pesscan_Ru_H_distance, pesscan_100_H2, pesscan_001_H2]:
name = f"dft_replay_{pesscan_job.name}"
settings = dft_settings(QEKPointsConfig(3, 3, 1)) + replay_settings(
pesscan_job.results.rkfpath()
)
replay_jobs[name] = plams.AMSJob(
settings=settings,
name=name,
)
ri = ResultsImporter.from_yaml(old_ref_dir)
for name, job in replay_jobs.items():
job.run()
ri.add_trajectory_singlepoints(job, properties=["energy", "forces"])
ri.store(new_ref_dir)
plot_pesscan(job)
[31.05|11:19:50] JOB dft_replay_PESScan_Ru_H_distance_m3gnet STARTED
[31.05|11:19:50] JOB dft_replay_PESScan_Ru_H_distance_m3gnet RUNNING
[31.05|11:36:42] JOB dft_replay_PESScan_Ru_H_distance_m3gnet FINISHED
[31.05|11:36:42] JOB dft_replay_PESScan_Ru_H_distance_m3gnet SUCCESSFUL
[31.05|11:36:43] JOB dft_replay_PESScan_100_H2_m3gnet STARTED
... output trimmed ....
[31.05|12:59:35] JOB dft_replay_PESScan_100_H2_m3gnet SUCCESSFUL
[31.05|12:59:35] JOB dft_replay_PESScan_001_H2_m3gnet STARTED
[31.05|12:59:35] JOB dft_replay_PESScan_001_H2_m3gnet RUNNING
[31.05|14:22:58] JOB dft_replay_PESScan_001_H2_m3gnet FINISHED
[31.05|14:22:58] JOB dft_replay_PESScan_001_H2_m3gnet SUCCESSFUL



See also¶
Python Script¶
#!/usr/bin/env python
# coding: utf-8
# ## Initial imports
import scm.plams as plams
from scm.params import ResultsImporter
from scm.plams import Settings, AMSJob, log, Molecule
from pathlib import Path
import matplotlib.pyplot as plt
# common_ru_h.py must exist in the current working directory
from common_ru_h import rotation, check_installation
# register dependencies for AMSjobs, to support submitting this notebook directly to a cluster in AMS2025+
# dependency: {} common_ru_h.py
# dependency: {} reference_data_1
# ## Initialize PLAMS working directory
old_ref_dir = "reference_data_1"
check_installation(old_ref_dir)
new_ref_dir = "reference_data_2"
ri = ResultsImporter.from_yaml(old_ref_dir)
plams.init()
# ## Load the optimized bulk Ru structure from the job collection
#
# The lattice was optimized in the previous notebook, and the structure was stored in the job collection.
#
# Let's retrieve it from the job collection and use it to construct Ru surface slabs.
optimized_bulk = ri.job_collection["hcp_lattopt_Ru_dft"].molecule
plams.plot_molecule(optimized_bulk, rotation=rotation)
plt.title("Bulk hcp Ru")
plt.gcf().savefig("picture1.png")
plt.clf()
# ## Cut out Ru(0001) and Ru(10-10) slabs
#
# The PLAMS Molecule class does not have any methods for creating slabs. However, we can use the ChemicalSystem in AMS2024+. The details can be found in the ``slice_slab()`` function in common_ru_h.py.
from common_ru_h import slice_slab
slab_100 = slice_slab(
optimized_bulk,
miller=(1, 0, 0),
thickness=7.0,
cell_z=15,
ref_atom=0,
)
slab_100 = slab_100.supercell(3, 2, 1)
for at in slab_100:
at.properties = Settings() # remove details about supercell generation
plams.plot_molecule(slab_100, rotation=rotation)
plt.title("Ru(10-10)")
plt.gcf().savefig("picture2.png")
plt.clf()
from common_ru_h import slice_slab
slab_001 = slice_slab(
optimized_bulk,
miller=(0, 0, 1),
thickness=7.0,
cell_z=15,
ref_atom=0,
)
slab_001 = slab_001.supercell([[3, 0, 0], [2, 4, 0], [0, 0, 1]])
for at in slab_001:
at.properties = Settings() # remove details about supercell generation
plams.plot_molecule(slab_001, rotation=rotation)
plt.title("Ru(0001)")
plt.gcf().savefig("picture3.png")
plt.clf()
# This slab is perhaps too thick, so let's remove the top atomic layer:
for at in slab_001:
if at.coords[2] > 10:
slab_001.delete_atom(at)
plams.plot_molecule(slab_001, rotation=rotation)
plt.title("Ru(0001)")
plt.gcf().savefig("picture4.png")
plt.clf()
# ## Add hydrogens
from common_ru_h import add_adsorbate
slab_100_H = add_adsorbate(slab_100, "H", frac_x=0, frac_y=0, delta_z=2.0)
# add second adsorbate, now delta_z=2.0 means 2 angstrom above the previous H
slab_100_H2 = add_adsorbate(slab_100_H, "H", frac_x=0.5, frac_y=0.5, delta_z=2.0)
plams.plot_molecule(slab_100_H2, rotation=rotation, radii=0.6)
plt.title("Ru(10-10) with H")
plt.gcf().savefig("picture5.png")
plt.clf()
from common_ru_h import add_adsorbate
slab_001_H = add_adsorbate(slab_001, "H", frac_x=0, frac_y=0, delta_z=2.0)
slab_001_H2 = add_adsorbate(slab_001_H, "H", frac_x=0.5, frac_y=0.5, delta_z=2.0)
plams.plot_molecule(slab_001_H2, rotation=rotation, radii=0.6)
plt.title("Ru(0001) with H")
plt.gcf().savefig("picture6.png")
plt.clf()
# ## Set up and run the jobs
# ### Ru-H distance bond scan on Ru(10-10)
from common_ru_h import get_surface_bond_scan_coordinates
from common_ru_h import (
get_bottom_atom_index,
pesscan_settings,
m3gnet_up_settings,
constraints_settings,
plot_pesscan,
)
system = slab_100_H
scan_coordinates = get_surface_bond_scan_coordinates(system, atom_index=len(system), start=1.2, end=2.5)
pesscan_Ru_H_distance = plams.AMSJob(
settings=(
pesscan_settings(scan_coordinates, n_points=6)
+ m3gnet_up_settings()
+ constraints_settings(get_bottom_atom_index(system))
),
name="PESScan_Ru_H_distance_m3gnet",
molecule=system,
)
pesscan_Ru_H_distance.run()
# !amsmovie {pesscan_Ru_H_distance.results.rkfpath()} # open in AMSmovie
plot_pesscan(pesscan_Ru_H_distance)
# ### Surface and bulk diffusion H on/in Ru(10-10)
#
# Note: These PES scans are just meant to sample H atoms diffusing through the slab and on the surface.
#
# You can also set up more intelligent PES scans (for example, to sample diffusion along specific pathways like hcp-bridge-fcc etc.). In these examples, the H atom simply diffuses from one corner of the cell to the opposite corner.
from common_ru_h import (
get_surface_diffusion_scan_coordinates,
get_bulk_diffusion_scan_coordinates,
)
from common_ru_h import (
get_bottom_atom_index,
pesscan_settings,
m3gnet_up_settings,
constraints_settings,
plot_pesscan,
)
system = slab_100_H2
scan_coordinates = get_surface_diffusion_scan_coordinates(system, atom_index=len(system) - 1)
scan_coordinates += get_bulk_diffusion_scan_coordinates(system, atom_index=len(system), delta_z=2.0)
pesscan_100_H2 = plams.AMSJob(
settings=(
pesscan_settings(scan_coordinates, n_points=20)
+ m3gnet_up_settings()
+ constraints_settings(get_bottom_atom_index(system))
),
name="PESScan_100_H2_m3gnet",
molecule=system,
)
pesscan_100_H2.run()
# !amsmovie {pesscan_100_H2.results.rkfpath()} # open in AMSmovie
plot_pesscan(pesscan_100_H2)
# ### Surface and bulk diffusion H on/in Ru(0001)
from common_ru_h import (
get_surface_diffusion_scan_coordinates,
get_bulk_diffusion_scan_coordinates,
)
from common_ru_h import (
get_bottom_atom_index,
pesscan_settings,
m3gnet_up_settings,
constraints_settings,
plot_pesscan,
)
system = slab_001_H2
scan_coordinates = get_surface_diffusion_scan_coordinates(system, atom_index=len(system) - 1)
scan_coordinates += get_bulk_diffusion_scan_coordinates(system, atom_index=len(system), delta_z=2.0)
pesscan_001_H2 = plams.AMSJob(
settings=(
pesscan_settings(scan_coordinates, n_points=20)
+ m3gnet_up_settings()
+ constraints_settings(get_bottom_atom_index(system))
),
name="PESScan_001_H2_m3gnet",
molecule=system,
)
pesscan_001_H2.run()
# !amsmovie {pesscan_001_H2.results.rkfpath()} # open in AMSmovie
plot_pesscan(pesscan_001_H2)
# ## Run DFT singlepoints on PES Scan structures and store on disk
from common_ru_h import dft_settings, replay_settings, QEKPointsConfig
replay_jobs = dict()
for pesscan_job in [pesscan_Ru_H_distance, pesscan_100_H2, pesscan_001_H2]:
name = f"dft_replay_{pesscan_job.name}"
settings = dft_settings(QEKPointsConfig(3, 3, 1)) + replay_settings(pesscan_job.results.rkfpath())
replay_jobs[name] = plams.AMSJob(
settings=settings,
name=name,
)
ri = ResultsImporter.from_yaml(old_ref_dir)
for name, job in replay_jobs.items():
job.run()
ri.add_trajectory_singlepoints(job, properties=["energy", "forces"])
ri.store(new_ref_dir)
plot_pesscan(job)