H-H Spin-Spin Coupling with ADF¶
See also the corresponding GUI tutorial.
This example shows how to set up an NMR calculation and extract basic results from it in Python. It follows the start of the GUI tutorial, which contains more background on the underlying workflow.
If you use the GUI, you can inspect the CPL and NMR program inputs through Details -> Run script. For spectrum analysis itself, the AMSspectra GUI module is usually the better tool; this scripting example focuses mainly on setting up and running the calculation.
Initial imports¶
from scm.base import ChemicalSystem
from scm.plams import AMSJob, Settings, view
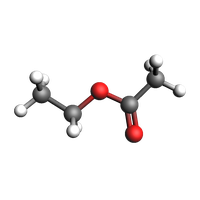
System¶
cs = ChemicalSystem(
"""
System
Atoms
O 1.48603879 -1.49561627 0.0
C 1.29751002 -0.30552432 0.0
O 0.07403584000000001 0.25228979 0.0
C -1.02449892 -0.67494471 0.0
C -2.30056502 0.12358769 0.0
C 2.36905363 0.74347075 0.0
H -0.9418758699999999 -1.31519741 0.88039373
H -0.9418758699999999 -1.31519741 -0.88039373
H -2.36617127 0.7582087199999999 0.88525259
H -3.15628689 -0.55419212 0.0
H -2.36617127 0.7582087199999999 -0.88525259
H 3.34355252 0.26293272 0.0
H 2.26362714 1.38098693 0.87932777
H 2.26362714 1.38098693 -0.87932777
End
End
"""
)
view(cs, guess_bonds=True, height=200, width=200)
Job settings: DFT and NMR¶
There are three calculations:
A DFT calculation with a main results file
adf.rkf. We must set theSave TAPE10input option. TheTAPE10file is later used by the$AMSBIN/nmrprogram.An
$AMSBIN/cplcalculation. This program modifiesadf.rkfAn
$AMSBIN/nmrcalculation. This program readsTAPE10and modifiesadf.rkf.
In the end, all results are stored on adf.rkf.
There is no PLAMS job for the $AMSBIN/cpl and $AMSBIN/nmr programs. Instead, we set up the calculation explicitly in the settings.runscript.postamble_lines option.
# DFT Settings
s = Settings()
s.input.ams.Task = "SinglePoint"
s.input.adf.Basis.Type = "TZ2P-J"
s.input.adf.Basis.Core = "None"
s.input.adf.Save = "TAPE10"
s.input.adf.xc.GGA = "OPBE"
s.input.adf.Symmetry = "NOSYM"
s.input.adf.NumericalQuality = "Good"
from typing import Mapping, List
def get_cpl_script(s: Mapping) -> List[str]:
ret = (
['"$AMSBIN/cpl" <<eor']
+ ["ADFFile adf.rkf"]
+ ["NMRCOUPLING"]
+ [f" {k} {v}" for k, v in s.items()]
+ ["END"]
+ ["eor"]
+ [""]
)
return ret
def get_nmr_script(s: Mapping) -> List[str]:
ret = (
['"$AMSBIN/nmr" <<eor']
+ ["ADFFile adf.rkf", "TAPE10File TAPE10"]
+ ["NMR"]
+ [f" {k} {v}" for k, v in s.items()]
+ ["End"]
+ ["eor"]
+ [""]
)
return ret
cpl_settings = dict(RespAllAtomsOfType="H", PertAllAtomsOfType="H", fc="", ADFGUI="")
nmr_settings = dict(out="", adfgui="", AllAtomsOfType="H", u1k="best", calc="all")
s.runscript.postamble_lines = get_cpl_script(cpl_settings) + get_nmr_script(nmr_settings)
Inspect the job¶
job = AMSJob(settings=s, molecule=cs, name="ethyl_acetate_nmr")
print(job.get_input()) # DFT settings in the normal input (ethyl_acetate_nmr.in)
Task SinglePoint
System
Atoms
O 1.48603879 -1.4956162699999997 0
C 1.29751002 -0.30552432 0
O 0.07403583999999999 0.25228979 0
C -1.02449892 -0.67494471 0
C -2.30056502 0.12358769 0
C 2.36905363 0.74347075 0
... output trimmed ....
End
NumericalQuality Good
Save TAPE10
Symmetry NOSYM
xc
GGA OPBE
End
EndEngine
print(job.get_runscript()) # post-processing utilities in the runscript
unset AMS_SWITCH_LOGFILE_AND_STDOUT
unset SCM_LOGFILE
AMS_JOBNAME="ethyl_acetate_nmr" AMS_RESULTSDIR=. $AMSBIN/ams --input="ethyl_acetate_nmr.in" < /dev/null
"$AMSBIN/cpl" <<eor
ADFFile adf.rkf
NMRCOUPLING
RespAllAtomsOfType H
PertAllAtomsOfType H
fc
... output trimmed ....
out
adfgui
AllAtomsOfType H
u1k best
calc all
End
eor
job.run();
[19.03|15:50:08] JOB ethyl_acetate_nmr STARTED
[19.03|15:50:08] JOB ethyl_acetate_nmr RUNNING
[19.03|15:58:01] JOB ethyl_acetate_nmr FINISHED
[19.03|15:58:01] JOB ethyl_acetate_nmr SUCCESSFUL
Results (basic)¶
Let’s see which NMR results are available on adf.rkf:
# job = AMSJob.load_external("/path/to/adf.rkf") # to load a previous job
props = job.results.get_engine_properties()
nmr_keys = [x for x in props.keys() if "NMR" in x]
print(nmr_keys)
['NMR Coupling J const InputOrder', 'NMR Shielding Tensor InputOrder', 'NMR Coupling K tens InputOrder', 'NMR Shieldings InputOrder', 'NMR Coupling K const InputOrder', 'NMR Coupling J tens InputOrder']
import numpy as np
import matplotlib.pyplot as plt
from scm.plams.tools.postprocess_results import broaden_results
sigma_ref_h = 31.7
shieldings = np.array(props["NMR Shieldings InputOrder"])
chemical_shifts = sigma_ref_h - np.array(shieldings)
x, y = broaden_results(
chemical_shifts,
areas=np.ones(chemical_shifts.shape), # weigh all peaks equally
broadening_width=0.01,
broadening_type="lorentzian",
x_data=(0, 5, 0.01), # x range and resolution
)
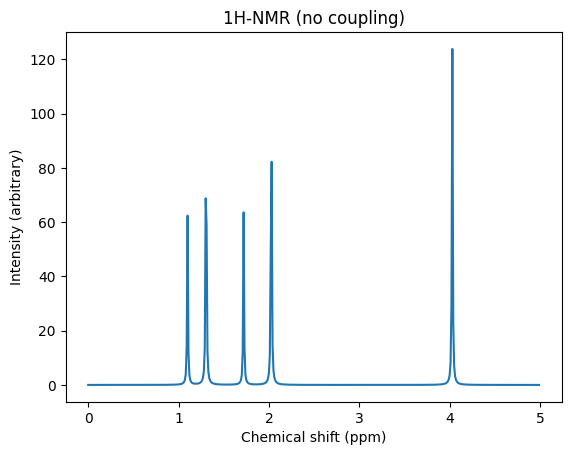
fig, ax = plt.subplots()
ax.set_title("1H-NMR (no coupling)")
ax.set_xlabel("Chemical shift (ppm)")
ax.set_ylabel("Intensity (arbitrary)")
ax.plot(x, y)
ax;
Conclusion¶
In this Python example you learned how to set up and run the NMR calculation.
For more detailed analysis, including multiplet splitting (coupling), see the GUI tutorial.
Recommendation: Use AMSspectra for further analysis.
See also¶
Python Script¶
#!/usr/bin/env python
# coding: utf-8
# ## Initial imports
from scm.base import ChemicalSystem
from scm.plams import AMSJob, Settings, view
# ## System
cs = ChemicalSystem(
"""
System
Atoms
O 1.48603879 -1.49561627 0.0
C 1.29751002 -0.30552432 0.0
O 0.07403584000000001 0.25228979 0.0
C -1.02449892 -0.67494471 0.0
C -2.30056502 0.12358769 0.0
C 2.36905363 0.74347075 0.0
H -0.9418758699999999 -1.31519741 0.88039373
H -0.9418758699999999 -1.31519741 -0.88039373
H -2.36617127 0.7582087199999999 0.88525259
H -3.15628689 -0.55419212 0.0
H -2.36617127 0.7582087199999999 -0.88525259
H 3.34355252 0.26293272 0.0
H 2.26362714 1.38098693 0.87932777
H 2.26362714 1.38098693 -0.87932777
End
End
"""
)
view(cs, guess_bonds=True, height=200, width=200, picture_path="picture1.png")
# ## Job settings: DFT and NMR
#
# There are three calculations:
#
# 1. A DFT calculation with a main results file ``adf.rkf``. We must set the ``Save TAPE10`` input option. The ``TAPE10`` file is later used by the ``$AMSBIN/nmr`` program.
#
# 2. An ``$AMSBIN/cpl`` calculation. This program *modifies* ``adf.rkf``
#
# 3. An ``$AMSBIN/nmr`` calculation. This program reads ``TAPE10`` and *modifies* ``adf.rkf``.
#
# In the end, all results are stored on ``adf.rkf``.
#
# There is no PLAMS job for the ``$AMSBIN/cpl`` and ``$AMSBIN/nmr`` programs. Instead, we set up the calculation explicitly in the ``settings.runscript.postamble_lines`` option.
# DFT Settings
s = Settings()
s.input.ams.Task = "SinglePoint"
s.input.adf.Basis.Type = "TZ2P-J"
s.input.adf.Basis.Core = "None"
s.input.adf.Save = "TAPE10"
s.input.adf.xc.GGA = "OPBE"
s.input.adf.Symmetry = "NOSYM"
s.input.adf.NumericalQuality = "Good"
from typing import Mapping, List
def get_cpl_script(s: Mapping) -> List[str]:
ret = (
['"$AMSBIN/cpl" <<eor']
+ ["ADFFile adf.rkf"]
+ ["NMRCOUPLING"]
+ [f" {k} {v}" for k, v in s.items()]
+ ["END"]
+ ["eor"]
+ [""]
)
return ret
def get_nmr_script(s: Mapping) -> List[str]:
ret = (
['"$AMSBIN/nmr" <<eor']
+ ["ADFFile adf.rkf", "TAPE10File TAPE10"]
+ ["NMR"]
+ [f" {k} {v}" for k, v in s.items()]
+ ["End"]
+ ["eor"]
+ [""]
)
return ret
cpl_settings = dict(RespAllAtomsOfType="H", PertAllAtomsOfType="H", fc="", ADFGUI="")
nmr_settings = dict(out="", adfgui="", AllAtomsOfType="H", u1k="best", calc="all")
s.runscript.postamble_lines = get_cpl_script(cpl_settings) + get_nmr_script(nmr_settings)
# ## Inspect the job
job = AMSJob(settings=s, molecule=cs, name="ethyl_acetate_nmr")
print(job.get_input()) # DFT settings in the normal input (ethyl_acetate_nmr.in)
print(job.get_runscript()) # post-processing utilities in the runscript
job.run()
# ## Results (basic)
#
# Let's see which NMR results are available on adf.rkf:
# job = AMSJob.load_external("/path/to/adf.rkf") # to load a previous job
props = job.results.get_engine_properties()
nmr_keys = [x for x in props.keys() if "NMR" in x]
print(nmr_keys)
import numpy as np
import matplotlib.pyplot as plt
from scm.plams.tools.postprocess_results import broaden_results
sigma_ref_h = 31.7
shieldings = np.array(props["NMR Shieldings InputOrder"])
chemical_shifts = sigma_ref_h - np.array(shieldings)
x, y = broaden_results(
chemical_shifts,
areas=np.ones(chemical_shifts.shape), # weigh all peaks equally
broadening_width=0.01,
broadening_type="lorentzian",
x_data=(0, 5, 0.01), # x range and resolution
)
fig, ax = plt.subplots()
ax.set_title("1H-NMR (no coupling)")
ax.set_xlabel("Chemical shift (ppm)")
ax.set_ylabel("Intensity (arbitrary)")
ax.plot(x, y)
ax
ax.figure.savefig("picture2.png")
# ## Conclusion
#
# In this Python example you learned how to set up and run the NMR calculation.
#
# For more detailed analysis, including multiplet splitting (coupling), see the GUI tutorial.
#
# **Recommendation**: Use AMSspectra for further analysis.