3.3. ReaxFF: Convert old file format¶
AMS2021 and earlier supported ReaxFF parametrizations using different file formats than ParAMS. This tutorial shows how to convert from the old formats to the new ParAMS format:
Old File |
+ |
(Optional File) |
= |
New File |
|---|---|---|---|---|
|
|
|
||
|
|
|||
|
|
|
Example files are located in $AMSHOME/scripting/scm/params/examples/import_old_ReaxFF.
3.3.1. Convert geo (and control) to job_collection.yaml¶
The old geo format distinguishes between:
single-point (
RUTYPE SINGLE POINT),geometry optimization (
RUTYPE MAXIT 100, where100is the maximum number of iterations), and“normal runs” (
RUTYPE NORMAL RUN). The meaning ofNORMAL RUNis defined in thecontrolfile.
The conversion automatically reads and writes:
The system charge (only one
MOLCHARGEline is allowed),The atomic positions, and
Bond and angle constraints (the force constants are approximately converted to the AMS format).
In the ParAMS GUI, select File → Open and browse to the geo file.
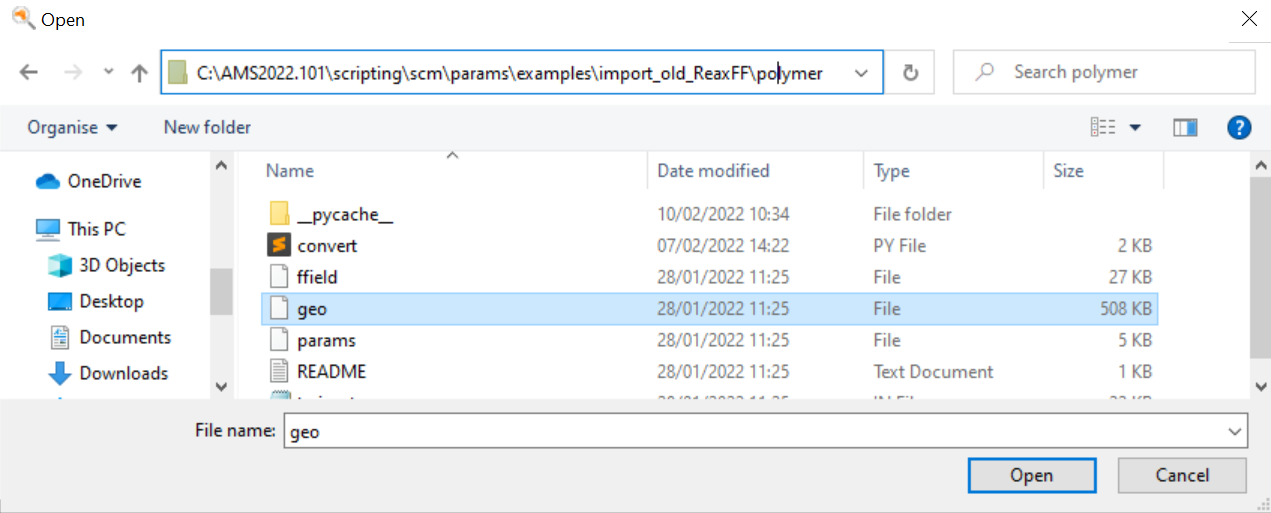
This adds the jobs to the job collection (and also reads
control, trainset.in, ffield and params if they are present in the same
directory).
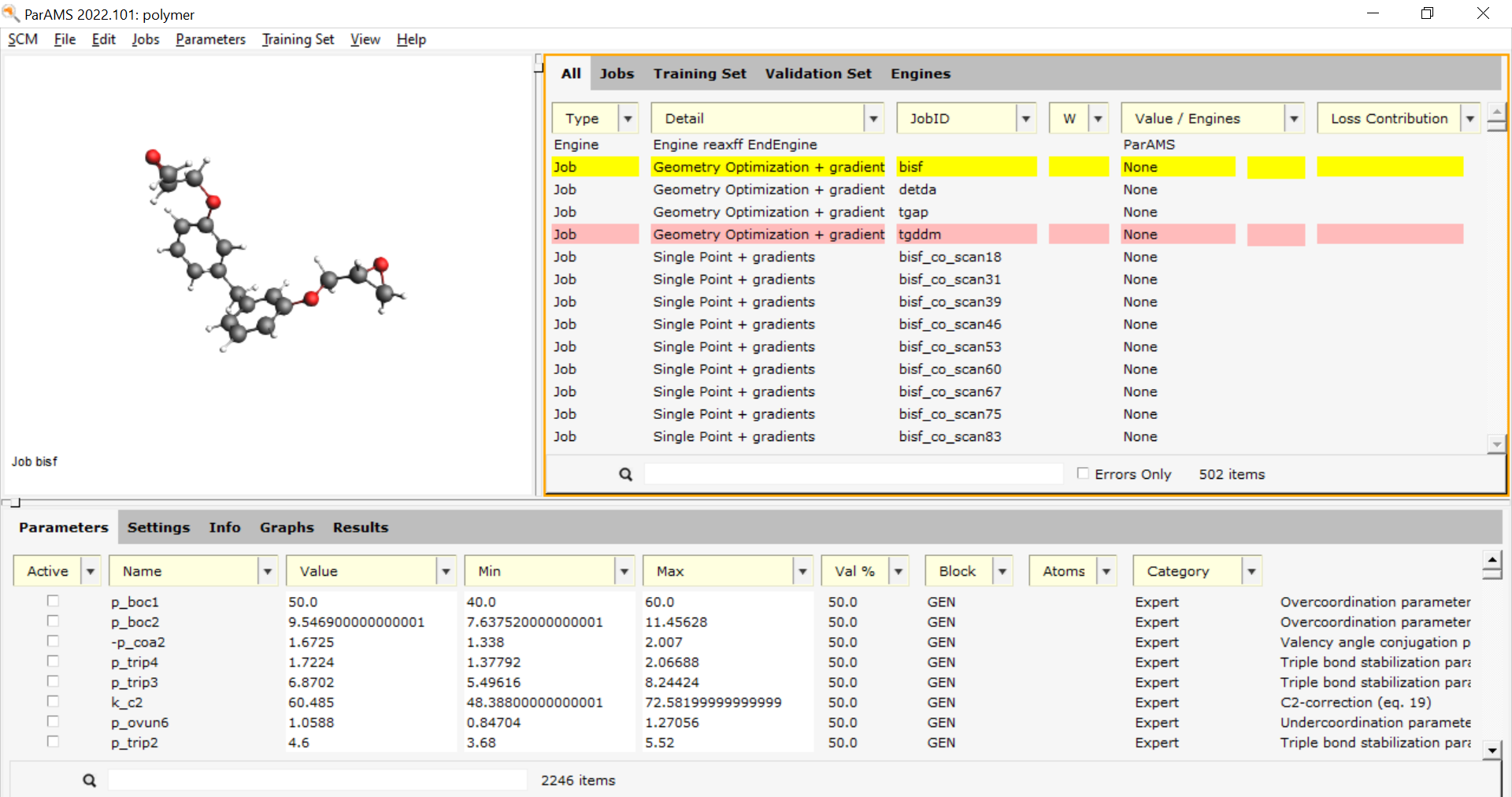
Important
When importing your own files:
If a control file is not found then any jobs with RUTYPE NORMAL RUN get converted to single point jobs.
To change them to geometry optimization, select the jobs you want to change on the Jobs panel, double-click in the Details column for one of them.
In the AMS Settings text box, type:
Task GeometryOptimization
GeometryOptimization
MaxIterations 30
PretendConverged Yes
End
Then click OK. See also the FAQ entry on MaxIterations and PretendConverged.
To convert to the job_collection.yaml format used by ParAMS, you need to specify the path to the geo file.
Optionally, you can also provide the path to the control file, or define the AMS settings for NORMAL RUN.
See the create_job_collection() in convert.py.
def create_job_collection(in_path: pathlib.Path, out_path: pathlib.Path):
"""Converts a 'geo' file to 'job_collection.yaml'.
Looks for a 'control' file to define the 'NORMAL RUN' run type. If not found,
'NORMAL RUN' defaults to a geometry optimization.
Parameters
----------
in_path
Path to directory containing the 'geo' and (optionally) 'control' file.
out_path
Path to directory in which to save 'job_collection.yaml'
"""
if (in_path / "control").exists():
print("'control' file being used for NORMAL RUN settings.")
normal_run_settings = str(in_path / "control")
else:
print("'control' file not found, using default settings for NORMAL RUN.")
normal_run_settings = Settings()
normal_run_settings.input.ams.Task = "GeometryOptimization"
normal_run_settings.input.ams.GeometryOptimization.MaxIterations = 10
# the PretendConverged option is used to make ParAMS transparently accept
# the last iteration of a geometry optimization if it does not converge within
# MaxIterations
normal_run_settings.input.ams.GeometryOptimization.PretendConverged = "Yes"
if (in_path / "geo").exists():
job_collection = geo_to_params(in_path / "geo", normal_run_settings=normal_run_settings)
job_collection.store(out_path / "job_collection.yaml")
else:
print("No 'geo' file found, 'job_collection.yaml' not built.")
3.3.2. Convert trainset.in to training_set.yaml¶
trainset.in file.This reads in the training set (and also reads
control, geo, ffield and params if they are present in the same
directory).
The old trainset.in format can easily be converted to training_set.yaml.
See the create_training_set() in convert.py.
def create_training_set(in_path: pathlib.Path, out_path: pathlib.Path):
"""Converts a 'trainset.in' file to 'training_set.yaml'.
Parameters
----------
in_path
Path to directory containing 'trainset.in'.
out_path
Path to directory in which to save 'training_set.yaml'
"""
if (in_path / "trainset.in").exists():
training_set = trainset_to_params(in_path / "trainset.in", use_weights=True)
training_set.store(out_path / "training_set.yaml")
# Extra summary of the training set, this is not used or needed by ParAMS, it is simply for user convenience.
with (out_path / "summary_training_set.txt").open("w") as f:
f.write("{:<8s} {:<8s} {:<19s} {}\n".format("Weight", "Sigma", "Reference_value", "Expression"))
for entry in training_set:
f.write(
"{:<8.3f} {:<8.3f} {:<8.3f} {:<10s} {}\n".format(
entry.weight, entry.sigma, entry.reference, entry.unit[0], entry.expression
)
)
else:
print("No 'trainset.in' file found, 'training_set.yaml' not built.")
use_weights=True(recommended, default) converts the “accuracy” from the oldtrainset.informat to a Weight in the newtraining_set.yamlformat.use_weights=Falsesets the Weight to 1.0 for all training set entries, and sets the Sigma to the “accuracy” fromtrainset.in.
For more information about Weight and Sigma, see Sigma vs. weight: What is the difference?.
3.3.3. Convert ffield (and params) to parameter_interface.yaml¶
The ffield file contains the force field parameters and their values.
The old params file (unrelated to ParAMS) contains the parameters to optimize and their allowed range of values.
params or ffield files.Alternatively:
This reads the parameter files (and also reads
control, trainset.in and geo if they are present in the same
directory).
To convert a ffield and params file to parameter_interface.yaml see the
create_parameter_interface() in convert.py.
def create_parameter_interface(in_path: pathlib.Path, out_path: pathlib.Path):
"""Converts a 'ffield' file to 'parameter_interface.yaml'.
Looks for a 'params' file to define the ranges and if a parameter is in/active. If not
found,all parameters are set inactive and the ranges are inferred from the magnitude
of the parameter value.
Parameters
----------
in_path
Path to directory containing the 'ffield' and (optionally) 'params' file.
out_path
Path to directory in which to save 'parameter_interface.yaml'
"""
if (in_path / "ffield").exists():
parameter_interface = ReaxFFParameters(in_path / "ffield")
if (in_path / "params").exists():
parameter_interface.read_paramsfile(in_path / "params")
else:
print("'params' file not found, ranges and 'active' set to default.")
parameter_interface.yaml_store(out_path / "parameter_interface.yaml")
else:
print("'ffield' file not found, 'parameter_interface.yaml' not built.")
3.3.4. Using convert.py¶
convert.py provides the stand-alone methods detailed above, but it can also be used as a script to bulk convert a directory of old files to the new format. The new files will be placed in a new directory called converted_to_params within the original directory.
"$AMSBIN/amspython" convert.py path/to/old/reaxff/files