Basic MD Trajectory Analysis with PLAMS¶
This example shows how to calculate several standard trajectory-analysis quantities from molecular dynamics:
the velocity autocorrelation function (VACF)
the diffusion coefficient from the integral of the VACF
the power spectrum from the Fourier transform of the VACF
the viscosity from the Green-Kubo relation using the off-diagonal pressure-tensor autocorrelation function
the density profile along the z axis
The example focuses on the mechanics of extracting these quantities. For production simulations, the trajectories must be much longer and the analysis must be checked for convergence. For a broader molecular-dynamics workflow example, see also Molecular Dynamics with Python.
Initial Imports¶
import os
import numpy as np
import matplotlib.pyplot as plt
from scm.plams import packmol, Settings, AMSJob, from_smiles, view
Run Simple MD Simulation of Water¶
Run a short MD simulation of 16 water molecules in a box.
mol = packmol(from_smiles("O"), n_molecules=16, density=1.0)
s = Settings()
s.input.ams.Task = "MolecularDynamics"
s.input.ReaxFF.ForceField = "Water2017.ff"
s.input.ams.MolecularDynamics.CalcPressure = "Yes"
s.input.ams.MolecularDynamics.InitialVelocities.Temperature = 300
s.input.ams.MolecularDynamics.Trajectory.SamplingFreq = 1
s.input.ams.MolecularDynamics.TimeStep = 0.5
s.input.ams.MolecularDynamics.NSteps = 2000
s.runscript.nproc = 1
os.environ["OMP_NUM_THREADS"] = "1"
job = AMSJob(settings=s, molecule=mol, name="md")
job.run();
[02.04|11:34:55] JOB md STARTED
[02.04|11:34:55] JOB md RUNNING
[02.04|11:35:02] JOB md FINISHED
[02.04|11:35:02] JOB md SUCCESSFUL
view(mol, direction="tilt_z", width=300, height=300, padding=-1)
Or alternatively, load a previously run MD job:
# job = AMSJob.load_external('/path/to/ams.rkf')
results = job.results
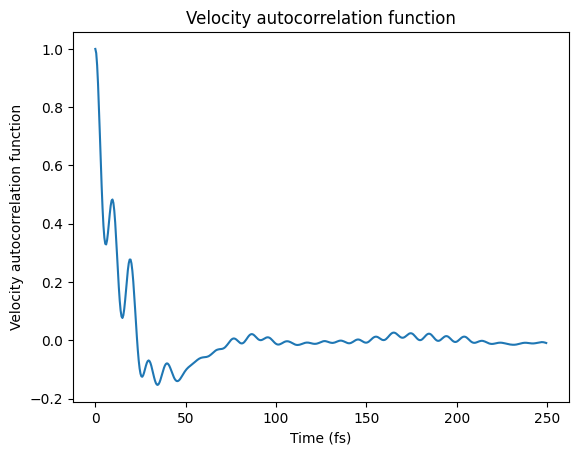
Velocity Autocorrelation Function¶
times, vacf = results.get_velocity_acf(start_fs=0, max_dt_fs=250, normalize=False)
normalized_vacf = vacf / vacf[0]
A = np.stack((times, normalized_vacf), axis=1)
np.savetxt("plams_vacf.txt", A, header="Time(fs) VACF")
fig, ax = plt.subplots()
ax.plot(times, normalized_vacf)
ax.set_xlabel("Time (fs)")
ax.set_ylabel("Velocity autocorrelation function")
ax.set_title("Velocity autocorrelation function")
ax;
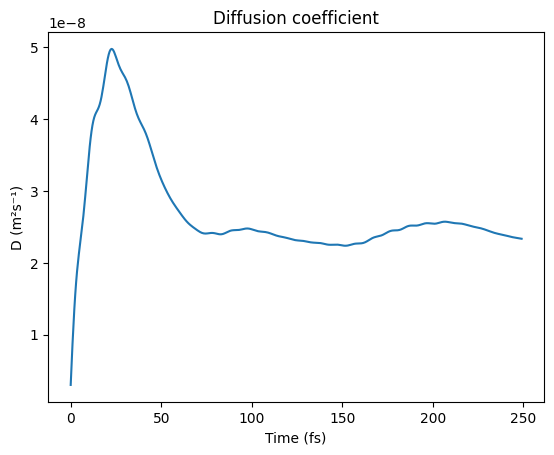
Diffusion Coefficient¶
t_D, D = results.get_diffusion_coefficient_from_velocity_acf(times, vacf)
A = np.stack((t_D, D), axis=1)
np.savetxt("plams_vacf_D.txt", A, header="time(fs) D(m^2*s^-1)")
fig, ax = plt.subplots()
ax.plot(t_D, D)
ax.set_xlabel("Time (fs)")
ax.set_ylabel("D (m²s⁻¹)")
ax.set_title("Diffusion coefficient")
ax;
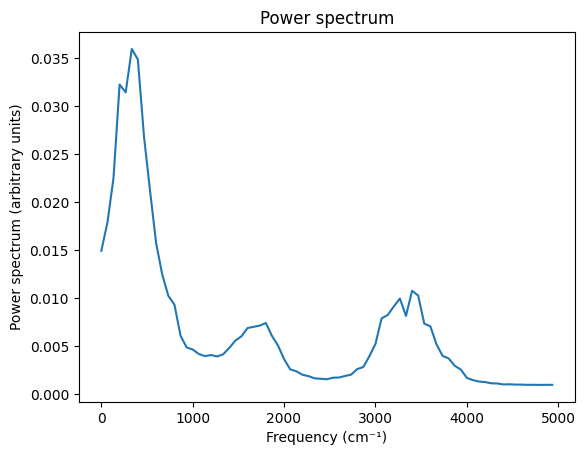
Power Spectrum¶
freq, intensities = results.get_power_spectrum(times, vacf, number_of_points=1000)
A = np.stack((freq, intensities), axis=1)
np.savetxt("plams_power_spectrum.txt", A, header="Frequency(cm^-1) PowerSpectrum")
fig, ax = plt.subplots()
ax.plot(freq, intensities)
ax.set_xlabel("Frequency (cm⁻¹)")
ax.set_ylabel("Power spectrum (arbitrary units)")
ax.set_title("Power spectrum")
ax;
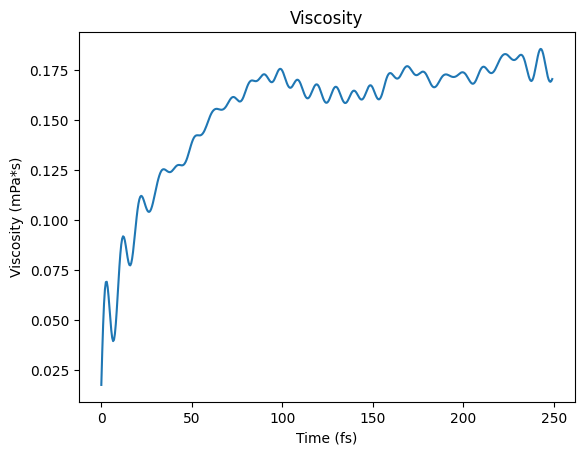
Green-Kubo Viscosity¶
t, viscosity = results.get_green_kubo_viscosity(
start_fs=0, max_dt_fs=250
) # do not do this for NPT simulations
A = np.stack((t, viscosity), axis=1)
np.savetxt("plams_green_kubo_viscosity.txt", A, header="Time(fs) Viscosity(mPa*s)")
fig, ax = plt.subplots()
ax.plot(t, viscosity)
ax.set_xlabel("Time (fs)")
ax.set_ylabel("Viscosity (mPa*s)")
ax.set_title("Viscosity")
ax;
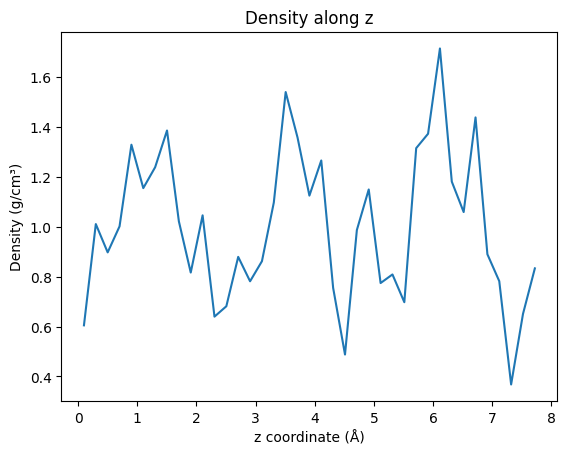
Density Along Axis¶
z, density = results.get_density_along_axis(
axis="z", density_type="mass", bin_width=0.2, atom_indices=None
)
A = np.stack((z, density), axis=1)
np.savetxt("plams_density_along_z.txt", A, header="z(angstrom) density(g/cm^3)")
fig, ax = plt.subplots()
ax.plot(z, density)
ax.set_xlabel("z coordinate (Å)")
ax.set_ylabel("Density (g/cm³)")
ax.set_title("Density along z")
ax;
See also¶
Python Script¶
#!/usr/bin/env python
# coding: utf-8
# ## Initial Imports
import os
import numpy as np
import matplotlib.pyplot as plt
from scm.plams import packmol, Settings, AMSJob, from_smiles, view
# ## Run Simple MD Simulation of Water
# Run a short MD simulation of 16 water molecules in a box.
mol = packmol(from_smiles("O"), n_molecules=16, density=1.0)
s = Settings()
s.input.ams.Task = "MolecularDynamics"
s.input.ReaxFF.ForceField = "Water2017.ff"
s.input.ams.MolecularDynamics.CalcPressure = "Yes"
s.input.ams.MolecularDynamics.InitialVelocities.Temperature = 300
s.input.ams.MolecularDynamics.Trajectory.SamplingFreq = 1
s.input.ams.MolecularDynamics.TimeStep = 0.5
s.input.ams.MolecularDynamics.NSteps = 2000
s.runscript.nproc = 1
os.environ["OMP_NUM_THREADS"] = "1"
job = AMSJob(settings=s, molecule=mol, name="md")
job.run()
view(mol, direction="tilt_z", width=300, height=300, padding=-1, picture_path="picture1.png")
# Or alternatively, load a previously run MD job:
# job = AMSJob.load_external('/path/to/ams.rkf')
results = job.results
# ## Velocity Autocorrelation Function
times, vacf = results.get_velocity_acf(start_fs=0, max_dt_fs=250, normalize=False)
normalized_vacf = vacf / vacf[0]
A = np.stack((times, normalized_vacf), axis=1)
np.savetxt("plams_vacf.txt", A, header="Time(fs) VACF")
fig, ax = plt.subplots()
ax.plot(times, normalized_vacf)
ax.set_xlabel("Time (fs)")
ax.set_ylabel("Velocity autocorrelation function")
ax.set_title("Velocity autocorrelation function")
ax
ax.figure.savefig("picture2.png")
# ## Diffusion Coefficient
t_D, D = results.get_diffusion_coefficient_from_velocity_acf(times, vacf)
A = np.stack((t_D, D), axis=1)
np.savetxt("plams_vacf_D.txt", A, header="time(fs) D(m^2*s^-1)")
fig, ax = plt.subplots()
ax.plot(t_D, D)
ax.set_xlabel("Time (fs)")
ax.set_ylabel("D (m²s⁻¹)")
ax.set_title("Diffusion coefficient")
ax
ax.figure.savefig("picture3.png")
# ## Power Spectrum
freq, intensities = results.get_power_spectrum(times, vacf, number_of_points=1000)
A = np.stack((freq, intensities), axis=1)
np.savetxt("plams_power_spectrum.txt", A, header="Frequency(cm^-1) PowerSpectrum")
fig, ax = plt.subplots()
ax.plot(freq, intensities)
ax.set_xlabel("Frequency (cm⁻¹)")
ax.set_ylabel("Power spectrum (arbitrary units)")
ax.set_title("Power spectrum")
ax
ax.figure.savefig("picture4.png")
# ## Green-Kubo Viscosity
t, viscosity = results.get_green_kubo_viscosity(start_fs=0, max_dt_fs=250) # do not do this for NPT simulations
A = np.stack((t, viscosity), axis=1)
np.savetxt("plams_green_kubo_viscosity.txt", A, header="Time(fs) Viscosity(mPa*s)")
fig, ax = plt.subplots()
ax.plot(t, viscosity)
ax.set_xlabel("Time (fs)")
ax.set_ylabel("Viscosity (mPa*s)")
ax.set_title("Viscosity")
ax
ax.figure.savefig("picture5.png")
# ## Density Along Axis
z, density = results.get_density_along_axis(axis="z", density_type="mass", bin_width=0.2, atom_indices=None)
A = np.stack((z, density), axis=1)
np.savetxt("plams_density_along_z.txt", A, header="z(angstrom) density(g/cm^3)")
fig, ax = plt.subplots()
ax.plot(z, density)
ax.set_xlabel("z coordinate (Å)")
ax.set_ylabel("Density (g/cm³)")
ax.set_title("Density along z")
ax
ax.figure.savefig("picture6.png")