Helium Dimer Dissociation Curve with ADF¶
Calculate energy-vs-distance for the helium dimer and compare the resulting energies point by point.
Initial Imports¶
import numpy as np
import pandas as pd
from scm.base import ChemicalSystem, Bond, Units
from scm.plams import Settings, AMSJob, view
Setup Dimer¶
We first create intial system of two Helium atoms at some minimum separation.
d_min = 2.2
d_max = 4.2
he_dimer = ChemicalSystem()
he_dimer.add_atom("He", coords=[0.0, 0.0, 0.0])
he_dimer.add_atom("He", coords=[d_min, 0.0, 0.0])
he_dimer.bonds.add_bond(0, 1, Bond(1.0))
print(he_dimer)
System
Atoms
He 0 0 0
He 2.2 0 0
End
BondOrders
1 2 1
End
End
Calculation Settings¶
We then set up the calculation settings. We will perform a PES scan over the He-He bond, taking 11 distances in our chosen range. For the engine we will use ADF with TZP/PBE+GrimmeD3.
settings = Settings()
settings.input.ams.task = "PESScan"
settings.input.ams.pesscan.scancoordinate.npoints = 11
settings.input.ams.pesscan.scancoordinate.distance = f"1 2 {d_min} {d_max}"
settings.input.adf.basis.type = "TZP"
settings.input.adf.xc.gga = "PBE"
settings.input.adf.xc.dispersion = "Grimme3"
Create and Run PESScan Job¶
The PES scan can now be run:
job = AMSJob(molecule=he_dimer, settings=settings)
job.run()
[09.04|17:55:44] JOB plamsjob STARTED
[09.04|17:55:44] JOB plamsjob RUNNING
[09.04|17:55:51] JOB plamsjob FINISHED
[09.04|17:55:51] JOB plamsjob SUCCESSFUL
<scm.plams.interfaces.adfsuite.ams.AMSResults at 0x12a396a60>
Results¶
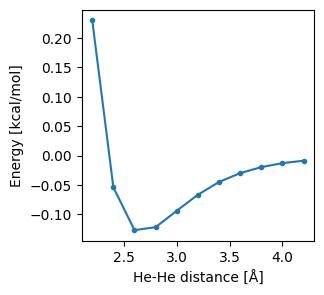
Results can be easily extracted using the get_pesscan_results method. We plot the energy as a function of the bond length, and the geometries either various points along the curve.
import pandas as pd
results = job.results.get_pesscan_results()
df = pd.DataFrame({"Distance": results["RaveledPESCoords"][0], "Energy": results["PES"]})
df["Distance"] *= Units.conversion_factor("Bohr", "Angstrom")
df["Energy"] *= Units.conversion_factor("Hartree", "kcal/mol")
df
Distance | Energy | |
|---|---|---|
0 | 2.2 | 0.230174 |
1 | 2.4 | -0.053810 |
2 | 2.6 | -0.126581 |
3 | 2.8 | -0.121580 |
4 | 3.0 | -0.093658 |
5 | 3.2 | -0.066451 |
6 | 3.4 | -0.044789 |
7 | 3.6 | -0.029741 |
8 | 3.8 | -0.019515 |
9 | 4.0 | -0.012803 |
10 | 4.2 | -0.008633 |
import matplotlib.pyplot as plt
fig, ax = plt.subplots(figsize=(3, 3))
ax.plot(df["Distance"], df["Energy"], ".-")
ax.set_xlabel("He-He distance [Å]")
ax.set_ylabel("Energy [kcal/mol]")
ax;
idx_min = df["Energy"].idxmin()
print(
f"Minimum energy {df['Energy'][idx_min]:.2f} kcal/mol at a separation of {df['Distance'][idx_min]:.1f}Å"
)
view(results["Molecules"][idx_min], height=150, width=150)
Minimum energy -0.13 kcal/mol at a separation of 2.6Å
[09.04|17:55:53] Starting Xvfb...
[09.04|17:55:53] Xvfb started
See also¶
Python Script¶
#!/usr/bin/env python
# coding: utf-8
# ## Initial Imports
import numpy as np
import pandas as pd
from scm.base import ChemicalSystem, Bond, Units
from scm.plams import Settings, AMSJob, view
# ## Setup Dimer
#
# We first create intial system of two Helium atoms at some minimum separation.
d_min = 2.2
d_max = 4.2
he_dimer = ChemicalSystem()
he_dimer.add_atom("He", coords=[0.0, 0.0, 0.0])
he_dimer.add_atom("He", coords=[d_min, 0.0, 0.0])
he_dimer.bonds.add_bond(0, 1, Bond(1.0))
print(he_dimer)
# ## Calculation Settings
#
# We then set up the calculation settings. We will perform a PES scan over the He-He bond, taking 11 distances in our chosen range. For the engine we will use ADF with TZP/PBE+GrimmeD3.
settings = Settings()
settings.input.ams.task = "PESScan"
settings.input.ams.pesscan.scancoordinate.npoints = 11
settings.input.ams.pesscan.scancoordinate.distance = f"1 2 {d_min} {d_max}"
settings.input.adf.basis.type = "TZP"
settings.input.adf.xc.gga = "PBE"
settings.input.adf.xc.dispersion = "Grimme3"
# ## Create and Run PESScan Job
#
# The PES scan can now be run:
job = AMSJob(molecule=he_dimer, settings=settings)
job.run()
# ## Results
#
# Results can be easily extracted using the `get_pesscan_results` method. We plot the energy as a function of the bond length, and the geometries either various points along the curve.
import pandas as pd
results = job.results.get_pesscan_results()
df = pd.DataFrame({"Distance": results["RaveledPESCoords"][0], "Energy": results["PES"]})
df["Distance"] *= Units.conversion_factor("Bohr", "Angstrom")
df["Energy"] *= Units.conversion_factor("Hartree", "kcal/mol")
print(df)
import matplotlib.pyplot as plt
fig, ax = plt.subplots(figsize=(3, 3))
ax.plot(df["Distance"], df["Energy"], ".-")
ax.set_xlabel("He-He distance [Å]")
ax.set_ylabel("Energy [kcal/mol]")
ax
ax.figure.savefig("picture1.png")
idx_min = df["Energy"].idxmin()
print(f"Minimum energy {df['Energy'][idx_min]:.2f} kcal/mol at a separation of {df['Distance'][idx_min]:.1f}Å")
view(results["Molecules"][idx_min], height=150, width=150, picture_path="picture2.png")