Diffusion Coefficient Supercell Dependence¶
Run Lennard-Jones argon simulations for several box sizes, calculate diffusion coefficients from the mean-squared displacement, and extrapolate the results to infinite system size with a linear fit in 1/L.
Diffusion coefficient supercell dependence¶
Finite-size effects make the diffusion coefficient depend on the size of the simulation cell. This example runs Lennard-Jones argon boxes of different sizes, computes the diffusion coefficient from the mean-squared displacement, and linearly extrapolates the results to the infinite-system limit.
from scm.plams import Settings
def get_lennard_jones_settings():
s = Settings()
s.input.lennardjones.eps = 3.785e-4
s.input.lennardjones.rmin = 3.81637
s.input.lennardjones.cutoff = 8.0
s.runscript.nproc = 1
return s
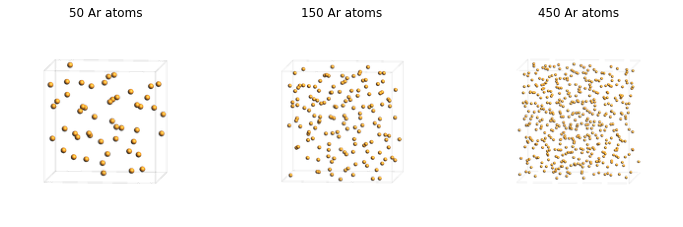
Set up three argon boxes at the same density and visualize the resulting supercells.
from scm.plams import Molecule, Atom, packmol, preoptimize, plot_image_grid, view
sizes = [50, 150, 450]
inverted_L_list = []
density = 1.0 # g/cm^3
Ar = Molecule()
Ar.add_atom(Atom(symbol="Ar", coords=(0, 0, 0)))
boxes = []
imgs = {}
for size in sizes:
mol = packmol(Ar, n_molecules=size, density=density)
mol = preoptimize(mol, settings=get_lennard_jones_settings(), maxiterations=50)
boxes.append(mol)
L = mol.lattice[0][0]
inverted_L_list.append(1.0 / L)
print(f"{size} Ar atoms, density = {density} g/cm^3")
imgs[f"{size} Ar atoms"] = view(mol, height=200, width=200, direction="tilt_z")
plot_image_grid(imgs, rows=1);
50 Ar atoms, density = 1.0 g/cm^3
150 Ar atoms, density = 1.0 g/cm^3
450 Ar atoms, density = 1.0 g/cm^3

For each system, run an NVT equilibration, an NVT production simulation, and an MSD analysis. The diffusion coefficients are then fitted as a function of 1/L.
D_list = []
temperature = 400
max_correlation_time_fs = 6000
start_time_fit_fs = 3000
from scm.plams import AMSNVTJob, AMSMSDJob, log
for size, mol in zip(sizes, boxes):
eq_job = AMSNVTJob(
name=f"eq_{size}",
settings=get_lennard_jones_settings(),
molecule=mol,
temperature=temperature,
nsteps=10000,
samplingfreq=500,
thermostat="Berendsen",
tau=100,
timestep=1.0,
writebonds=False,
writecharges=False,
writemolecules=False,
writevelocities=False,
)
eq_job.run()
prod_job = AMSNVTJob.restart_from(
eq_job,
name=f"prod_{size}",
nsteps=60000,
samplingfreq=25,
temperature=temperature,
thermostat="NHC",
)
prod_job.run()
msd_job = AMSMSDJob(
prod_job, name=f"msd_{size}", max_correlation_time_fs=max_correlation_time_fs
)
msd_job.run()
D = msd_job.results.get_diffusion_coefficient(start_time_fit_fs=start_time_fit_fs)
D_list.append(D)
log(f"{size} atoms: D = {D:.2e} m^2 s^-1")
[23.03|16:01:20] JOB eq_50 STARTED
[23.03|16:01:20] JOB eq_50 RUNNING
[23.03|16:01:22] JOB eq_50 FINISHED
[23.03|16:01:22] JOB eq_50 SUCCESSFUL
[23.03|16:01:22] JOB prod_50 STARTED
[23.03|16:01:22] JOB prod_50 RUNNING
[23.03|16:01:34] JOB prod_50 FINISHED
[23.03|16:01:34] JOB prod_50 SUCCESSFUL
[23.03|16:01:34] JOB msd_50 STARTED
[23.03|16:01:34] JOB msd_50 RUNNING
... output trimmed ....
[23.03|16:02:21] JOB eq_450 SUCCESSFUL
[23.03|16:02:21] JOB prod_450 STARTED
[23.03|16:02:21] JOB prod_450 RUNNING
[23.03|16:03:40] JOB prod_450 FINISHED
[23.03|16:03:41] JOB prod_450 SUCCESSFUL
[23.03|16:03:41] JOB msd_450 STARTED
[23.03|16:03:41] JOB msd_450 RUNNING
[23.03|16:03:42] JOB msd_450 FINISHED
[23.03|16:03:42] JOB msd_450 SUCCESSFUL
[23.03|16:03:42] 450 atoms: D = 2.44e-08 m^2 s^-1
import matplotlib.pyplot as plt
from scm.plams import log
from scm.plams.tools.plot import linear_fit_extrapolate_to_0
fit_x, fit_y, slope, intercept = linear_fit_extrapolate_to_0(inverted_L_list, D_list)
dilute_D = intercept
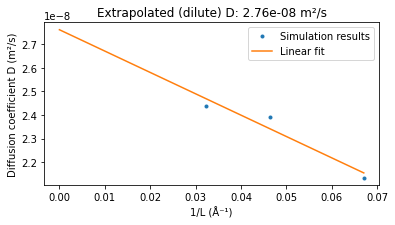
title = f"Extrapolated (dilute) D: {dilute_D:.2e} m²/s"
log(title)
fig, ax = plt.subplots(figsize=(6, 3))
ax.plot(inverted_L_list, D_list, ".", label="Simulation results")
ax.plot(fit_x, fit_y, label="Linear fit")
ax.set_title(title)
ax.legend()
ax.set_xlabel("1/L (Å⁻¹)")
ax.set_ylabel("Diffusion coefficient D (m²/s)")
ax;
[23.03|16:03:42] Extrapolated (dilute) D: 2.76e-08 m²/s
See also¶
Python Script¶
#!/usr/bin/env python
# coding: utf-8
# ## Diffusion coefficient supercell dependence
#
# Finite-size effects make the diffusion coefficient depend on the size of the simulation cell. This example runs Lennard-Jones argon boxes of different sizes, computes the diffusion coefficient from the mean-squared displacement, and linearly extrapolates the results to the infinite-system limit.
#
from scm.plams import Settings
def get_lennard_jones_settings():
s = Settings()
s.input.lennardjones.eps = 3.785e-4
s.input.lennardjones.rmin = 3.81637
s.input.lennardjones.cutoff = 8.0
s.runscript.nproc = 1
return s
# Set up three argon boxes at the same density and visualize the resulting supercells.
from scm.plams import Molecule, Atom, packmol, preoptimize, plot_image_grid, view
sizes = [50, 150, 450]
inverted_L_list = []
density = 1.0 # g/cm^3
Ar = Molecule()
Ar.add_atom(Atom(symbol="Ar", coords=(0, 0, 0)))
boxes = []
imgs = {}
for size in sizes:
mol = packmol(Ar, n_molecules=size, density=density)
mol = preoptimize(mol, settings=get_lennard_jones_settings(), maxiterations=50)
boxes.append(mol)
L = mol.lattice[0][0]
inverted_L_list.append(1.0 / L)
print(f"{size} Ar atoms, density = {density} g/cm^3")
imgs[f"{size} Ar atoms"] = view(mol, height=200, width=200, direction="tilt_z")
plot_image_grid(imgs, rows=1)
# For each system, run an NVT equilibration, an NVT production simulation, and an MSD analysis. The diffusion coefficients are then fitted as a function of ``1/L``.
#
D_list = []
temperature = 400
max_correlation_time_fs = 6000
start_time_fit_fs = 3000
from scm.plams import AMSNVTJob, AMSMSDJob, log
for size, mol in zip(sizes, boxes):
eq_job = AMSNVTJob(
name=f"eq_{size}",
settings=get_lennard_jones_settings(),
molecule=mol,
temperature=temperature,
nsteps=10000,
samplingfreq=500,
thermostat="Berendsen",
tau=100,
timestep=1.0,
writebonds=False,
writecharges=False,
writemolecules=False,
writevelocities=False,
)
eq_job.run()
prod_job = AMSNVTJob.restart_from(
eq_job,
name=f"prod_{size}",
nsteps=60000,
samplingfreq=25,
temperature=temperature,
thermostat="NHC",
)
prod_job.run()
msd_job = AMSMSDJob(prod_job, name=f"msd_{size}", max_correlation_time_fs=max_correlation_time_fs)
msd_job.run()
D = msd_job.results.get_diffusion_coefficient(start_time_fit_fs=start_time_fit_fs)
D_list.append(D)
log(f"{size} atoms: D = {D:.2e} m^2 s^-1")
import matplotlib.pyplot as plt
from scm.plams import log
from scm.plams.tools.plot import linear_fit_extrapolate_to_0
fit_x, fit_y, slope, intercept = linear_fit_extrapolate_to_0(inverted_L_list, D_list)
dilute_D = intercept
title = f"Extrapolated (dilute) D: {dilute_D:.2e} m²/s"
log(title)
fig, ax = plt.subplots(figsize=(6, 3))
ax.plot(inverted_L_list, D_list, ".", label="Simulation results")
ax.plot(fit_x, fit_y, label="Linear fit")
ax.set_title(title)
ax.legend()
ax.set_xlabel("1/L (Å⁻¹)")
ax.set_ylabel("Diffusion coefficient D (m²/s)")
ax
ax.figure.savefig("picture1.png")