Molecule Tools for Inspection and Manipulation¶
Explore ChemicalSystem methods for creating subsystems, bond handling, alignment, and structure manipulation.
Molecule Tools¶
Get a fragment and add H atoms¶
import scm.plams as plams
from scm.base import ChemicalSystem
system = ChemicalSystem.from_smiles("NC(CO)OCC=O")
plams.view(system, width=200, height=200)
[10.04|11:49:35] Starting Xvfb...
[10.04|11:49:35] Xvfb started
Get a fragment from the initial system.
fragment = system.extract_atoms([0, 1, 2, 3, 4, 5])
plams.view(fragment, direction="along_pca3", width=200, height=200)
print(fragment)
System
Atoms
N -0.8067647372863869 -1.6846900332599775 -0.2529030289460961
C -0.5865921942423814 -0.2897182936531656 0.1385551093669827
C -1.8520160304405069 0.5420466773061617 -0.10225001764394374
O -1.6383896887149183 1.8593832808319926 0.3233284897788621
O 0.47220924648540097 0.25958681517678306 -0.6254688148278047
C 1.7192462670365973 -0.12916614709198904 -0.08751486466412597
End
BondOrders
1 2 1
2 3 1
2 5 1
3 4 1
5 6 1
End
End
Add H atoms to fill the missing bonds
fragment_h = fragment.copy()
fragment_h.add_hydrogen_atoms()
plams.view(fragment_h, direction="along_pca3", width=200, height=200)
import matplotlib.pyplot as plt
import numpy as np
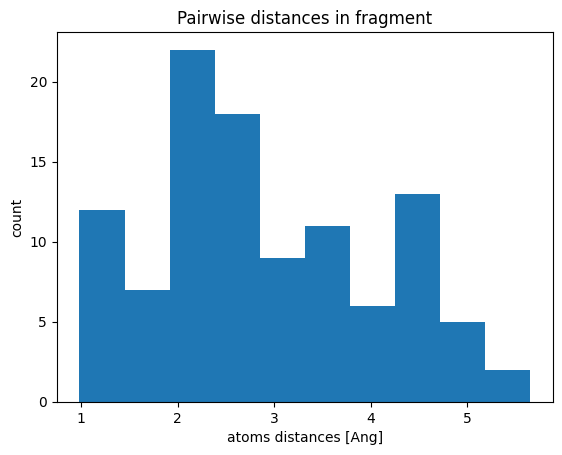
matrix = fragment_h.distance_matrix()
mask = np.triu(np.ones_like(matrix, dtype=bool), k=1)
fig, ax = plt.subplots()
ax.hist(matrix[mask])
ax.set_title("Pairwise distances in fragment")
ax.set_xlabel("atoms distances [Ang]")
ax.set_ylabel("count")
ax;
Visualize systems¶
With plams.view you can render ChemicalSystem objects directly. You can also compose multiple rendered images with plams.plot_image_grid.
system = ChemicalSystem.from_smiles("NC(CO)OCC=O")
small_system = ChemicalSystem.from_smiles("NC")
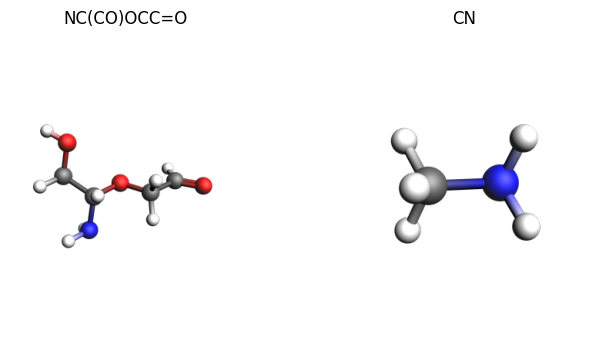
images = {
system.to_smiles(): plams.view(system, width=300),
small_system.to_smiles(): plams.view(small_system, width=300),
}
plams.plot_image_grid(images);
To visualize with RDKit as 2D images:
plams.plot_grid_molecules([system, small_system])
See also¶
Python Script¶
#!/usr/bin/env python
# coding: utf-8
# ## Molecule Tools
# ### Get a fragment and add H atoms
import scm.plams as plams
from scm.base import ChemicalSystem
system = ChemicalSystem.from_smiles("NC(CO)OCC=O")
plams.view(system, width=200, height=200, picture_path="picture1.png")
# Get a fragment from the initial system.
fragment = system.extract_atoms([0, 1, 2, 3, 4, 5])
plams.view(fragment, direction="along_pca3", width=200, height=200, picture_path="picture2.png")
print(fragment)
# Add H atoms to fill the missing bonds
fragment_h = fragment.copy()
fragment_h.add_hydrogen_atoms()
plams.view(fragment_h, direction="along_pca3", width=200, height=200, picture_path="picture3.png")
import matplotlib.pyplot as plt
import numpy as np
matrix = fragment_h.distance_matrix()
mask = np.triu(np.ones_like(matrix, dtype=bool), k=1)
fig, ax = plt.subplots()
ax.hist(matrix[mask])
ax.set_title("Pairwise distances in fragment")
ax.set_xlabel("atoms distances [Ang]")
ax.set_ylabel("count")
ax
ax.figure.savefig("picture4.png")
# ### Visualize systems
# With ``plams.view`` you can render ChemicalSystem objects directly. You can also compose multiple rendered images with `plams.plot_image_grid`.
system = ChemicalSystem.from_smiles("NC(CO)OCC=O")
small_system = ChemicalSystem.from_smiles("NC")
images = {
system.to_smiles(): plams.view(system, width=300),
small_system.to_smiles(): plams.view(small_system, width=300),
}
plams.plot_image_grid(images, save_path="picture5.png")
# To visualize with RDKit as 2D images:
plams.plot_grid_molecules([system, small_system])